Actin
Topic: Biology
 From HandWiki - Reading time: 53 min
From HandWiki - Reading time: 53 min
| Actin | |||||||||
|---|---|---|---|---|---|---|---|---|---|
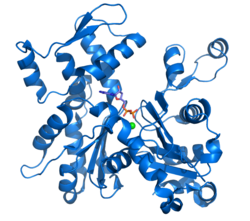 Ribbon diagram of G-actin. ADP bound to actin's active site (multi color sticks near center of figure) as well as a complexed calcium dication (green sphere) are highlighted.[1] | |||||||||
| Identifiers | |||||||||
| Symbol | Actin | ||||||||
| Pfam | PF00022 | ||||||||
| InterPro | IPR004000 | ||||||||
| PROSITE | PDOC00340 | ||||||||
| SCOP2 | 2btf / SCOPe / SUPFAM | ||||||||
| |||||||||
Actin is a family of globular multi-functional proteins that form microfilaments in the cytoskeleton, and the thin filaments in muscle fibrils. It is found in essentially all eukaryotic cells, where it may be present at a concentration of over 100 μM; its mass is roughly 42 kDa, with a diameter of 4 to 7 nm.
An actin protein is the monomeric subunit of two types of filaments in cells: microfilaments, one of the three major components of the cytoskeleton, and thin filaments, part of the contractile apparatus in muscle cells. It can be present as either a free monomer called G-actin (globular) or as part of a linear polymer microfilament called F-actin (filamentous), both of which are essential for such important cellular functions as the mobility and contraction of cells during cell division.
Actin participates in many important cellular processes, including muscle contraction, cell motility, cell division and cytokinesis, vesicle and organelle movement, cell signaling, and the establishment and maintenance of cell junctions and cell shape. Many of these processes are mediated by extensive and intimate interactions of actin with cellular membranes.[2] In vertebrates, three main groups of actin isoforms, alpha, beta, and gamma have been identified. The alpha actins, found in muscle tissues, are a major constituent of the contractile apparatus. The beta and gamma actins coexist in most cell types as components of the cytoskeleton, and as mediators of internal cell motility. It is believed that the diverse range of structures formed by actin enabling it to fulfill such a large range of functions is regulated through the binding of tropomyosin along the filaments.[3]
A cell's ability to dynamically form microfilaments provides the scaffolding that allows it to rapidly remodel itself in response to its environment or to the organism's internal signals, for example, to increase cell membrane absorption or increase cell adhesion in order to form cell tissue. Other enzymes or organelles such as cilia can be anchored to this scaffolding in order to control the deformation of the external cell membrane, which allows endocytosis and cytokinesis. It can also produce movement either by itself or with the help of molecular motors. Actin therefore contributes to processes such as the intracellular transport of vesicles and organelles as well as muscular contraction and cellular migration. It therefore plays an important role in embryogenesis, the healing of wounds, and the invasivity of cancer cells. The evolutionary origin of actin can be traced to prokaryotic cells, which have equivalent proteins.[4] Actin homologs from prokaryotes and archaea polymerize into different helical or linear filaments consisting of one or multiple strands. However the in-strand contacts and nucleotide binding sites are preserved in prokaryotes and in archaea.[5] Lastly, actin plays an important role in the control of gene expression.
A large number of illnesses and diseases are caused by mutations in alleles of the genes that regulate the production of actin or of its associated proteins. The production of actin is also key to the process of infection by some pathogenic microorganisms. Mutations in the different genes that regulate actin production in humans can cause muscular diseases, variations in the size and function of the heart as well as deafness. The make-up of the cytoskeleton is also related to the pathogenicity of intracellular bacteria and viruses, particularly in the processes related to evading the actions of the immune system.[6]
Function
Actin's primary role in the cell is to form linear polymers called microfilaments that serve various functions in the cell's structure, trafficking networks, migration, and replication.[7] The multifaceted role of actin relies on a few of the microfilaments' properties: First, the formation of actin filaments is reversible, and their function often involves undergoing rapid polymerization and depolymerization. Second, microfilaments are polarized – i.e. the two ends of a filament are distinct from one another. Third, actin filaments can bind to many other proteins, which together help modify and organize microfilaments for their diverse functions.[7]
In most cells actin filaments form larger-scale networks which are essential for many key functions:[8]
- Actin networks give mechanical support to cells and provide trafficking routes through the cytoplasm to aid signal transduction.
- Rapid assembly and disassembly of actin network enables cells to migrate (Cell migration).
Actin is extremely abundant in most cells, comprising 1–5% of the total protein mass of most cells, and 10% of muscle cells.[7]
The actin protein is found in both the cytoplasm and the cell nucleus.[9] Its location is regulated by cell membrane signal transduction pathways that integrate the stimuli that a cell receives stimulating the restructuring of the actin networks in response.[10]
Cytoskeleton
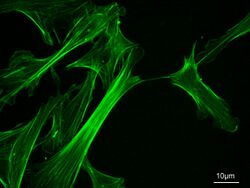
There are a number of different types of actin with slightly different structures and functions. α-actin is found exclusively in muscle fibres, while β- and γ-actin are found in other cells. As the latter types have a high turnover rate the majority of them are found outside permanent structures. Microfilaments found in cells other than muscle cells are present in three forms:[11]
- Microfilament networks - Animal cells commonly have a cell cortex under the cell membrane that contains a large number of microfilaments, which precludes the presence of organelles. This network is connected with numerous receptors that relay signals to the outside of a cell.
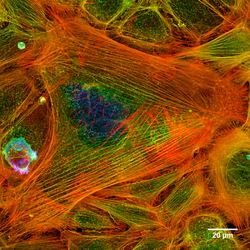
- Periodic actin rings - A periodic structure constructed of evenly spaced actin rings is found in axons.[12] In this structure, the actin rings, together with spectrin tetramers that link the neighboring actin rings, form a cohesive cytoskeleton that supports the axon membrane. The structure periodicity may also regulate the sodium ion channels in axons.
Yeasts
Actin's cytoskeleton is key to the processes of endocytosis, cytokinesis, determination of cell polarity and morphogenesis in yeasts. In addition to relying on actin, these processes involve 20 or 30 associated proteins, which all have a high degree of evolutionary conservation, along with many signalling molecules. Together these elements allow a spatially and temporally modulated assembly that defines a cell's response to both internal and external stimuli.[13]
Yeasts contain three main elements that are associated with actin: patches, cables, and rings. Despite not being present for long, these structures are subject to a dynamic equilibrium due to continual polymerization and depolymerization. They possess a number of accessory proteins including ADF/cofilin, which has a molecular weight of 16kDa and is coded for by a single gene, called COF1; Aip1, a cofilin cofactor that promotes the disassembly of microfilaments; Srv2/CAP, a process regulator related to adenylate cyclase proteins; a profilin with a molecular weight of approximately 14 kDa that is related/associated with actin monomers; and twinfilin, a 40 kDa protein involved in the organization of patches.[13]
Plants
Plant genome studies have revealed the existence of protein isovariants within the actin family of genes. Within Arabidopsis thaliana, a model organism, there are ten types of actin, six profilins, and dozens of myosins. This diversity is explained by the evolutionary necessity of possessing variants that slightly differ in their temporal and spatial expression.[4] The majority of these proteins were jointly expressed in the tissue analysed. Actin networks are distributed throughout the cytoplasm of cells that have been cultivated in vitro. There is a concentration of the network around the nucleus that is connected via spokes to the cellular cortex, this network is highly dynamic, with a continuous polymerization and depolymerization.[14]

Even though the majority of plant cells have a cell wall that defines their morphology, their microfilaments can generate sufficient force to achieve a number of cellular activities, such as the cytoplasmic currents generated by the microfilaments and myosin. Actin is also involved in the movement of organelles and in cellular morphogenesis, which involve cell division as well as the elongation and differentiation of the cell.[16]
The most notable proteins associated with the actin cytoskeleton in plants include:[16] villin, which belongs to the same family as gelsolin/severin and is able to cut microfilaments and bind actin monomers in the presence of calcium cations; fimbrin, which is able to recognize and unite actin monomers and which is involved in the formation of networks (by a different regulation process from that of animals and yeasts);[17] formins, which are able to act as an F-actin polymerization nucleating agent; myosin, a typical molecular motor that is specific to eukaryotes and which in Arabidopsis thaliana is coded for by 17 genes in two distinct classes; CHUP1, which can bind actin and is implicated in the spatial distribution of chloroplasts in the cell; KAM1/MUR3 that define the morphology of the Golgi apparatus as well as the composition of xyloglucans in the cell wall; NtWLIM1, which facilitates the emergence of actin cell structures; and ERD10, which is involved in the association of organelles within membranes and microfilaments and which seems to play a role that is involved in an organism's reaction to stress.
Nuclear actin
Nuclear actin was first noticed and described in 1977 by Clark and Merriam.[18] Authors describe a protein present in the nuclear fraction, obtained from Xenopus laevis oocytes, which shows the same features as skeletal muscle actin. Since that time there have been many scientific reports about the structure and functions of actin in the nucleus (for review see: Hofmann 2009.[19]) The controlled level of actin in the nucleus, its interaction with actin-binding proteins (ABP) and the presence of different isoforms allows actin to play an important role in many important nuclear processes.[20]
Transport through the nuclear membrane
The actin sequence does not contain a nuclear localization signal. The small size of actin (about 43 kDa) allows it to enter the nucleus by passive diffusion.[21] The import of actin into the nucleus (probably in a complex with cofilin) is facilitated by the import protein importin 9.[22]
Low levels of actin in the nucleus seems to be important, because actin has two nuclear export signals (NES) in its sequence. Microinjected actin is quickly removed from the nucleus to the cytoplasm. Actin is exported at least in two ways, through exportin 1 and exportin 6.[23][24] Specific modifications, such as SUMOylation, allows for nuclear actin retention. A mutation preventing SUMOylation causes rapid export of beta actin from the nucleus.[25]
Organization
Nuclear actin exists mainly as a monomer, but can also form dynamic oligomers and short polymers.[26][27][28] Nuclear actin organization varies in different cell types. For example, in Xenopus oocytes (with higher nuclear actin level in comparison to somatic cells) actin forms filaments, which stabilize nucleus architecture. These filaments can be observed under the microscope thanks to fluorophore-conjugated phalloidin staining.[18][21]
In somatic cell nuclei, however, actin filaments cannot be observed using this technique.[29] The DNase I inhibition assay, the only test which allows the quantification of the polymerized actin directly in biological samples, has revealed that endogenous nuclear actin indeed occurs mainly in a monomeric form.[28]
Precisely controlled level of actin in the cell nucleus, lower than in the cytoplasm, prevents the formation of filaments. The polymerization is also reduced by the limited access to actin monomers, which are bound in complexes with ABPs, mainly cofilin.[30]
Actin isoforms
Different isoforms of actin are present in the cell nucleus. The level of actin isoforms may change in response to stimulation of cell growth or arrest of proliferation and transcriptional activity.[31] Research on nuclear actin is focused on isoform beta.[32][33][34][35] However the use of antibodies directed against different actin isoforms allows identifying not only the cytoplasmic beta in the cell nucleus, but also alpha- and gamma-actin in certain cell types.[28][36][37] The presence of different isoforms of actin may have a significant effect on its function in nuclear processes, as the level of individual isoforms can be controlled independently.[28]
Functions
Functions of actin in the nucleus are associated with its ability to polymerize and interact with various ABPs and with structural elements of the nucleus. Nuclear actin is involved in:
- Architecture of the nucleus - Interaction of actin with alpha II-spectrin and other proteins are important for maintaining proper shape of the nucleus.[38][39]
- Transcription – Actin is involved in chromatin reorganization,[9][32][40][41] transcription initiation and interaction with the transcription complex.[42] Actin takes part in the regulation of chromatin structure,[43][44][45] interacting with RNA polymerase I,[35] II[33] and III.[34] In Pol I transcription, actin and myosin (MYO1C, which binds DNA) act as a molecular motor. For Pol II transcription, β-actin is needed for the formation of the preinitiation complex. Pol III contains β-actin as a subunit. Actin can also be a component of chromatin remodelling complexes as well as pre-mRNP particles (that is, precursor messenger RNA bundled in proteins), and is involved in nuclear export of RNAs and proteins.[46]
- Regulation of gene activity – Actin binds to the regulatory regions of different kinds of genes.[47][48][49][50] Actin's ability to regulate gene activity is used in the molecular reprogramming method, which allows differentiated cells return to their embryonic state.[49][51]
- Translocation of the activated chromosome fragment from under membrane region to euchromatin where transcription starts. This movement requires the interaction of actin and myosin.[52][53]
- Integration of different cellular compartments. Actin is a molecule that integrates cytoplasmic and nuclear signal transduction pathways.[54] An example is the activation of transcription in response to serum stimulation of cells in vitro.[55][56][57]
- Immune response - Nuclear actin polymerizes upon T-cell receptor stimulation and is required for cytokine expression and antibody production in vivo.[58]
Due to its ability to undergo conformational changes and interaction with many proteins, actin acts as a regulator of formation and activity of protein complexes such as transcriptional complex.[42]
Cell movement
Actin is also involved in cell movement. A meshwork of actin filaments marks the forward edge of a moving cell, and the polymerization of new actin filaments pushes the cell membrane forward in protrusions called lamellipodia.[59] These membrane protrusions then attach to the substrate, forming structures known as focal adhesions that connect to the actin network.[59] Once attached, the rear of the cell body contracts squeezing its contents forward past the adhesion point.[59] Once the adhesion point has moved to the rear of the cell, the cell disassembles it, allowing the rear of the cell to move forward.[59]
Actin/myosin movement
In addition to the physical force generated by actin polymerization, microfilaments facilitate the movement of various intracellular components by serving as the roadway along which a family of motor proteins called myosins travel.[60]
Muscle contraction
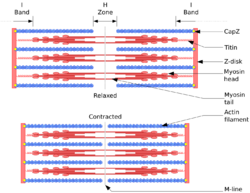
Actin plays a particularly prominent role in muscle cells, which consist largely of repeated bundles of actin and myosin II.[61] Each repeated unit – called a sarcomere – consists of two sets of oppositely oriented F-actin strands ("thin filaments"), interlaced with bundles of myosin ("thick filaments"). The two sets of actin strands are oriented with their (+) ends embedded in either end of the sarcomere in delimiting structures called Z-disks.[61] The myosin fibrils are in the middle between the sets of actin filaments, with strands facing in both directions. When the muscle contracts, the myosin threads move along the actin filaments towards the (+) end, pulling the ends of the sarcomere together and shortening it by around 70% of its length.[61] In order to move along the actin thread, myosin must hydrolyze ATP; thus ATP serves as the energy source for muscle contraction.[61]
At times of rest, the proteins tropomyosin and troponin bind to the actin filaments, preventing the attachment of myosin.[61] When an activation signal (i.e. an action potential) arrives at the muscle fiber, it triggers the release of Ca2+ from the sarcoplasmic reticulum into the cytosol. The resulting spike in cytosolic calcium rapidly releases tropomyosin and troponin from the actin thread, allowing myosin to bind, and muscle contracation to begin.[62]
Cell division
In the final stages of cell division, many cells form a ring of actin at the cell's midpoint. This ring, aptly called the "contractile ring", uses a similar mechanism as muscle fibers where myosin II pulls along the actin ring, causing it to contract.[63] This contraction cleaves the parent cell into two, completing cytokinesis.[63] The contractile ring is composed of actin, myosin, anillin, and α-actinin.[64] In the fission yeast Schizosaccharomyces pombe, actin is actively formed in the constricting ring with the participation of Arp3, the formin Cdc12, profilin, and WASp, along with preformed microfilaments. Once the ring has been constructed the structure is maintained by a continual assembly and disassembly that, aided by the Arp2/3 complex and formins, is key to one of the central processes of cytokinesis.[65]
Intracellular trafficking
Actin-myosin pairs can also participate in the trafficking of various membrane vesicles and organelles within the cell. Myosin V is activated by binding to various cargo receptors on organelles, and then moves along an actin filament towards the (+) end, pulling its cargo along with it.[66]
These nonconventional myosins use ATP hydrolysis to transport cargo, such as vesicles and organelles, in a directed fashion much faster than diffusion. Myosin V walks towards the barbed end of actin filaments, while myosin VI walks toward the pointed end. Most actin filaments are arranged with the barbed end toward the cellular membrane and the pointed end toward the cellular interior. This arrangement allows myosin V to be an effective motor for the export of cargos, and myosin VI to be an effective motor for import.
Other biological processes
File:Diverse-roles-of-actin-in-C.-elegans-early-embryogenesis-1471-213X-7-142-S9.ogv
The traditional image of actin's function relates it to the maintenance of the cytoskeleton and, therefore, the organization and movement of organelles, as well as the determination of a cell's shape.[11] However, actin has a wider role in eukaryotic cell physiology, in addition to similar functions in prokaryotes.
- Apoptosis. During programmed cell death the ICE/ced-3 family of proteases (one of the interleukin-1β-converter proteases) degrade actin into two fragments in vivo; one of the fragments is 15 kDa and the other 31 kDa. This represents one of the mechanisms involved in destroying cell viability that form the basis of apoptosis.[67] The protease calpain has also been shown to be involved in this type of cell destruction;[68] just as the use of calpain inhibitors has been shown to decrease actin proteolysis and the degradation of DNA (another of the characteristic elements of apoptosis).[69] On the other hand, the stress-induced triggering of apoptosis causes the reorganization of the actin cytoskeleton (which also involves its polymerization), giving rise to structures called stress fibers; this is activated by the MAP kinase pathway.[70]

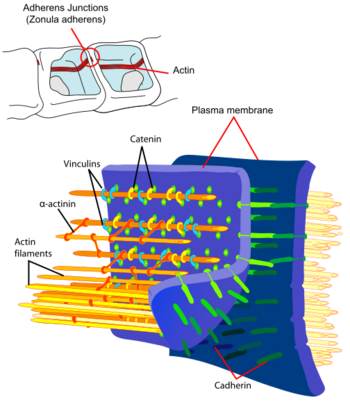
- Cellular adhesion and development. The adhesion between cells is a characteristic of multicellular organisms that enables tissue specialization and therefore increases cell complexity. Adhesion of cell epithelia involves the actin cytoskeleton in each of the joined cells as well as cadherins acting as extracellular elements with the connection between the two mediated by catenins.[71] Interfering in actin dynamics has repercussions for an organism's development, in fact actin is such a crucial element that systems of redundant genes are available. For example, if the α-actinin or gelation factor gene has been removed in Dictyostelium individuals do not show an anomalous phenotype possibly due to the fact that each of the proteins can perform the function of the other. However, the development of double mutations that lack both gene types is affected.[72]
- Gene expression modulation. Actin's state of polymerization affects the pattern of gene expression. In 1997, it was discovered that cytocalasin D-mediated depolymerization in Schwann cells causes a specific pattern of expression for the genes involved in the myelinization of this type of nerve cell.[73] F-actin has been shown to modify the transcriptome in some of the life stages of unicellular organisms, such as the fungus Candida albicans.[74] In addition, proteins that are similar to actin play a regulatory role during spermatogenesis in mice[75] and, in yeasts, actin-like proteins are thought to play a role in the regulation of gene expression.[76] In fact, actin is capable of acting as a transcription initiator when it reacts with a type of nuclear myosin that interacts with RNA polymerases and other enzymes involved in the transcription process.[9]
- Stereocilia dynamics. Some cells develop fine filliform outgrowths on their surface that have a mechanosensory function. For example, this type of organelle is present in the Organ of Corti, which is located in the ear. The main characteristic of these structures is that their length can be modified.[77] The molecular architecture of the stereocilia includes a paracrystalline actin core in dynamic equilibrium with the monomers present in the adjacent cytosol. Type VI and VIIa myosins are present throughout this core, while myosin XVa is present in its extremities in quantities that are proportional to the length of the stereocilia.[78]
- Intrinsic chirality. Actomyosin networks have been implicated in generating an intrinsic chirality in individual cells.[79] Cells grown out on chiral surfaces can show a directional left/right bias that is actomyosin dependent.[80][81]
Structure
Cellular actin has two forms: monomeric globules called G-actin and polymeric filaments called F-actin or microfilaments. Each molecule of actin is bound to a molecule of adenosine triphosphate (ATP) or adenosine diphosphate (ADP) that is associated with a Mg2+ cation. The most commonly found forms of actin, compared to all the possible combinations, are ATP-G-Actin and ADP-F-actin.[82][83]
G-actin

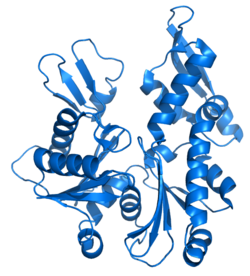
Monomeric actin, or G-actin, has a globular structure consisting of two lobes separated by a deep cleft.[84] The bottom of the cleft represents the “ATPase fold”, a structure conserved among ATP and GTP-binding proteins that binds to a magnesium ion and a molecule of ATP.[84] Binding of ATP or ADP is required to stabilize each actin monomer; without one of these molecules bound, actin quickly becomes denatured.[84]
The X-ray crystallography model of actin that was produced by Kabsch from the striated muscle tissue of rabbits is the most commonly used in structural studies as it was the first to be purified. The G-actin crystallized by Kabsch is approximately 67 x 40 x 37 Å in size, has a molecular mass of 41,785 Da and an estimated isoelectric point of 4.8. Its net charge at pH = 7 is -7.[85][86]
- Primary structure
Elzinga and co-workers first determined the complete peptide sequence for this type of actin in 1973, with later work by the same author adding further detail to the model. It contains 374 amino acid residues. Its N-terminus is highly acidic and starts with an acetyled aspartate in its amino group. While its C-terminus is alkaline and is formed by a phenylalanine preceded by a cysteine, which has a degree of functional importance. Both extremes are in close proximity within the I-subdomain. An anomalous Nτ-methylhistidine is located at position 73.[86]
- Tertiary structure — domains
The tertiary structure is formed by two domains known as the large and the small, which are separated by a cleft centred around the location of the bond with ATP-ADP+Pi. Below this there is a deeper notch called a “groove”. In the native state, despite their names, both have a comparable depth.[85]
The normal convention in topological studies means that a protein is shown with the biggest domain on the left-hand side and the smallest domain on the right-hand side. In this position the smaller domain is in turn divided into two: subdomain I (lower position, residues 1–32, 70–144, and 338–374) and subdomain II (upper position, residues 33–69). The larger domain is also divided in two: subdomain III (lower, residues 145–180 and 270–337) and subdomain IV (higher, residues 181–269). The exposed areas of subdomains I and III are referred to as the “barbed” ends, while the exposed areas of domains II and IV are termed the “pointed" ends. This nomenclature refers to the fact that, due to the small mass of subdomain II actin is polar; the importance of this will be discussed below in the discussion on assembly dynamics. Some authors call the subdomains Ia, Ib, IIa, and IIb, respectively.[87]
- Other important structures
The most notable supersecondary structure is a five chain beta sheet that is composed of a β-meander and a β-α-β clockwise unit. It is present in both domains suggesting that the protein arose from gene duplication.[88]
- The adenosine nucleotide binding site is located between two beta hairpin-shaped structures pertaining to the I and III domains. The residues that are involved are Asp11-Lys18 and Asp154-His161 respectively.
- The divalent cation binding site is located just below that for the adenosine nucleotide. In vivo it is most often formed by Mg2+ or Ca2+ while in vitro it is formed by a chelating structure made up of Lys18 and two oxygens from the nucleotide's α-and β-phosphates. This calcium is coordinated with six water molecules that are retained by the amino acids Asp11, Asp154, and Gln137. They form a complex with the nucleotide that restricts the movements of the so-called "hinge" region, located between residues 137 and 144. This maintains the native form of the protein until its withdrawal denatures the actin monomer. This region is also important because it determines whether the protein's cleft is in the "open" or "closed" conformation.[1][87]
- It is highly likely that there are at least three other centres with a lesser affinity (intermediate) and still others with a low affinity for divalent cations. It has been suggested that these centres may play a role in the polymerization of actin by acting during the activation stage.[87]
- There is a structure in subdomain 2 that is called the “D-loop” because it binds with DNase I, it is located between the His40 and Gly48 residues. It has the appearance of a disorderly element in the majority of crystals, but it looks like a β-sheet when it is complexed with DNase I. It has been proposed that the key event in polymerization is probably the propagation of a conformational change from the centre of the bond with the nucleotide to this domain, which changes from a loop to a spiral.[1] However, this hypothesis has been refuted by other studies.[89]
F-actin

Under various conditions, G-actin molecules polymerize into longer threads called "filamentous-" or "F-actin". These F-actin threads are typically composed of two helical strands of actin wound around each other, forming a 7 to 9 nanometer wide helix that repeats every 72 nanometers (or every 14 G-actin subunits).[91] In F-actin threads, G-actin molecules are all oriented in the same direction. The two ends of the F-actin thread are distinct from one another. At one end – designated the (−) end – the ATP-binding cleft of the terminal actin molecule is facing outward. At the opposite end – designated (+) – the ATP-binding cleft is buried in the filament, contacting the neighboring actin molecule.[91] As F-actin threads grow, new molecules tend to join at the (+) end of an existing F-actin strand. Conversely, threads tend to shrink by shedding actin monomers from the strand's (-) end.[91]
Some proteins, such as cofilin appear to increase the angle of turn, but again this could be interpreted as the establishment of different structural states. These could be important in the polymerization process.[92]
There is less agreement regarding measurements of the turn radius and filament thickness: while the first models assigned a length of 25 Å, current X-ray diffraction data, backed up by cryo-electron microscopy suggests a length of 23.7 Å. These studies have shown the precise contact points between monomers. Some are formed with units of the same chain, between the "barbed" end on one monomer and the "pointed" end of the next one. While the monomers in adjacent chains make lateral contact through projections from subdomain IV, with the most important projections being those formed by the C-terminus and the hydrophobic link formed by three bodies involving residues 39–42, 201–203, and 286. This model suggests that a filament is formed by monomers in a "sheet" formation, in which the subdomains turn about themselves, this form is also found in the bacterial actin homologue MreB.[93]
The terms "pointed" and "barbed" referring to the two ends of the microfilaments derive from their appearance under transmission electron microscopy when samples are examined following a preparation technique called "decoration". This method consists of the addition of myosin S1 fragments to tissue that has been fixed with tannic acid. This myosin forms polar bonds with actin monomers, giving rise to a configuration that looks like arrows with feather fletchings along its shaft, where the shaft is the actin and the fletchings are the myosin. Following this logic, the end of the microfilament that does not have any protruding myosin is called the point of the arrow (- end) and the other end is called the barbed end (+ end).[94] A S1 fragment is composed of the head and neck domains of myosin II. Under physiological conditions, G-actin (the monomer form) is transformed to F-actin (the polymer form) by ATP, where the role of ATP is essential.[95]
The helical F-actin filament found in muscles also contains a tropomyosin molecule, which is a 40 nanometre long protein that is wrapped around the F-actin helix.[96] During the resting phase the tropomyosin covers the actin's active sites so that the actin-myosin interaction cannot take place and produce muscular contraction. There are other protein molecules bound to the tropomyosin thread, these are the troponins that have three polymers: troponin I, troponin T, and troponin C.[97]
F-actin is both strong and dynamic. Unlike other polymers, such as DNA, whose constituent elements are bound together with covalent bonds, the monomers of actin filaments are assembled by weaker bonds.[98] The lateral bonds with neighbouring monomers resolve this anomaly, which in theory should weaken the structure as they can be broken by thermal agitation. In addition, the weak bonds give the advantage that the filament ends can easily release or incorporate monomers. This means that the filaments can be rapidly remodelled and can change cellular structure in response to an environmental stimulus. Which, along with the biochemical mechanism by which it is brought about is known as the "assembly dynamic".[6]
Folding

Actin can spontaneously acquire a large part of its tertiary structure.[100] However, the way it acquires its fully functional form from its newly synthesized native form is special and almost unique in protein chemistry. The reason for this special route could be the need to avoid the presence of incorrectly folded actin monomers, which could be toxic as they can act as inefficient polymerization terminators. Nevertheless, it is key to establishing the stability of the cytoskeleton, and additionally, it is an essential process for coordinating the cell cycle.[101][102]
CCT is required in order to ensure that folding takes place correctly. CCT is a group II chaperonin, a large protein complex that assists in the folding of other proteins. CCT is formed of a double ring of eight different subunits (hetero-octameric) and it differs from group I chaperonins like GroEL, which is found in Eubacteria and in eukaryotic organelles, as it does not require a co-chaperone to act as a lid over the central catalytic cavity. Substrates bind to CCT through specific domains. It was initially thought that it only bound with actin and tubulin, although recent immunoprecipitation studies have shown that it interacts with a large number of polypeptides, which possibly function as substrates. It acts through ATP-dependent conformational changes that on occasion require several rounds of liberation and catalysis in order to complete a reaction.[103]
In order to successfully complete their folding, both actin and tubulin need to interact with another protein called prefoldin, which is a heterohexameric complex (formed by six distinct subunits), in an interaction that is so specific that the molecules have coevolved[citation needed]. Actin complexes with prefoldin while it is still being formed, when it is approximately 145 amino acids long, specifically those at the N-terminal.[104]
Different recognition sub-units are used for actin or tubulin although there is some overlap. In actin the subunits that bind with prefoldin are probably PFD3 and PFD4, which bind in two places one between residues 60–79 and the other between residues 170–198. The actin is recognized, loaded, and delivered to the cytosolic chaperonin (CCT) in an open conformation by the inner end of prefoldin's "tentacles” (see the image and note).[100] The contact when actin is delivered is so brief that a tertiary complex is not formed, immediately freeing the prefoldin.[99]

The CCT then causes actin's sequential folding by forming bonds with its subunits rather than simply enclosing it in its cavity.[105] This is why it possesses specific recognition areas in its apical β-domain. The first stage in the folding consists of the recognition of residues 245–249. Next, other determinants establish contact.[106] Both actin and tubulin bind to CCT in open conformations in the absence of ATP. In actin's case, two subunits are bound during each conformational change, whereas for tubulin binding takes place with four subunits. Actin has specific binding sequences, which interact with the δ and β-CCT subunits or with δ-CCT and ε-CCT. After AMP-PNP is bound to CCT the substrates move within the chaperonin's cavity. It also seems that in the case of actin, the CAP protein is required as a possible cofactor in actin's final folding states.[102]
The exact manner by which this process is regulated is still not fully understood, but it is known that the protein PhLP3 (a protein similar to phosducin) inhibits its activity through the formation of a tertiary complex.[103]
ATPase’s catalytic mechanism
Actin is an ATPase, which means that it is an enzyme that hydrolyzes ATP. This group of enzymes is characterised by their slow reaction rates. It is known that this ATPase is “active”, that is, its speed increases by some 40,000 times when the actin forms part of a filament.[92] A reference value for this rate of hydrolysis under ideal conditions is around 0.3 s−1. Then, the Pi remains bound to the actin next to the ADP for a long time, until it is cooperatively liberated from the interior of the filament.[107][108]
The exact molecular details of the catalytic mechanism are still not fully understood. Although there is much debate on this issue, it seems certain that a "closed" conformation is required for the hydrolysis of ATP, and it is thought that the residues that are involved in the process move to the appropriate distance.[92] The glutamic acid Glu137 is one of the key residues, which is located in subdomain 1. Its function is to bind the water molecule that produces a nucleophilic attack on the ATP's γ-phosphate bond, while the nucleotide is strongly bound to subdomains 3 and 4. The slowness of the catalytic process is due to the large distance and skewed position of the water molecule in relation to the reactant. It is highly likely that the conformational change produced by the rotation of the domains between actin's G and F forms moves the Glu137 closer allowing its hydrolysis. This model suggests that the polymerization and ATPase's function would be decoupled straight away.[93][96] The "open" to "closed" transformation between G and F forms and its implications on the relative motion of several key residues and the formation of water wires have been characterized in molecular dynamics and QM/MM simulations.[109][110]
Assembly dynamics
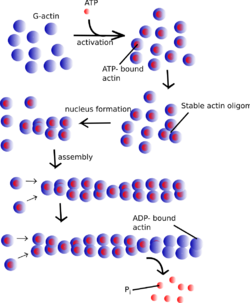

Actin filaments are often rapidly assembled and disassembled, allowing them to generate force and support cell movement.[111] Assembly classically occurs in three steps. First, the "nucleation phase", in which two to three G-actin molecules slowly join to form a small oligomer that will nucleate further growth. Second, the "elongation phase", when the actin filament rapidly grows by the addition of many actin molecules to both ends. As the filament grows, actin molecules are added to the (+) end of the filament around 10 times faster than to the (−) end, and so filaments tend to primarily grow at the (+) end.[112] Third, the "steady-state phase", where an equillibrium is reached as actin molecules join and leave the filament at the same rate, maintaining the filament's length.[111] While the filament's length remains constant in the steady-state phase, new molecules are constantly being added to the (+) end and falling off the (−) end, a phenomenon called "treadmilling" as a given actin molecule would appear to move along the strand.[113] In isolation, whether a filament will grow or shrink, and how quickly, are determined by the concentration of G-actin around the filament;[112] however, in cells, the dynamics of actin filaments are heavily influenced by various actin-binding proteins.
Actin binding proteins
The actin cytoskeleton in vivo is not exclusively composed of actin, other proteins are required for its formation, continuance, and function. These proteins are called actin-binding proteins and they are involved in actin's polymerization, depolymerization, stability, and organisation.[114] The diversity of these proteins is such that actin is thought to be the protein that takes part in the greatest number of protein-protein interactions.[115]
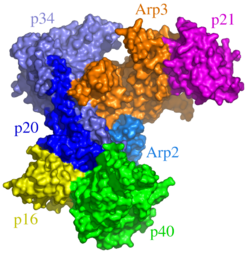
The nucleation of new actin filaments – the rate-limiting step in actin polymerization – is aided by actin-nucleating proteins such as formins and the Arp2/3 complex.[117] Formins help to nucleate long actin filaments. They bind two free actin-ATP molecules, bringing them together. Then as the filament begins to grow, formin moves along the (+) end of the growing filament, all the while recruiting actin-binding proteins that promote filament growth, and excluding capping proteins that would block filament extension.[117] Branches in actin filaments are typically nucleated by the Arp2/3 complex in concert with nucleation promoting factors. Nucleation promoting factors bind two free G-actin molecules, then recruit and activate the Arp2/3 complex. The activated Arp2/3 complex attaches to an existing actin filament, and uses the two bound G-actin molecules to nucleate a new actin filament branching off of the old one at a 70° angle.[118]
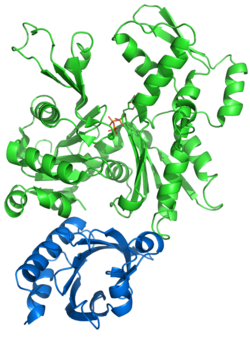
As filaments grow, the pool of available G-actin molecules is managed by G-actin-binding proteins such as profilin and thymosin β-4. Profilin ensures a supply of available actin-ATP by binding to ADP-bound G-actin and promoting the exchange of ADP for ATP. Profilin's binding to the actin molecule physically blocks its addition to a filament's (−) end, but permits it to join the (+) end. Once the actin-ATP has joined the filament, profilin releases it.[113] As formins promote the nucleation and extension of new actin filaments, they recruit profilin to the area, increasing the local concentration of actin-ATP to boost filament growth.[117] In contrast, thymosin β-4 binds and sequesters actin-ATP, preventing it from joining a microfilament.[120]
Once an actin fiber is established, the dynamics of its growth or collapse are influenced by numerous proteins. Existing strands can be interrupted by filament cleaving proteins, such as cofilin and gelsolin. Cofilin binds along two actin-ADP molecules in a filament, forcing a movement that destabilizes the filament and causes it to break.[121] Gelsolin inserts itself between actin molecules in a filament, disrupting the filament. After the filament breaks, gelsolin remains attached to the new (+) end, preventing it from growing, thus forcing its disassembly.[120]
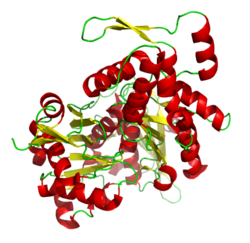
Other proteins bind to the ends of actin filaments, stabilizing them. These are called "capping proteins" and include CapZ and tropomodulin. CapZ binds the (+) end of a filament, preventing further addition or loss of actin from that end.[120] Tropomodulin binds to a filament's (−) end, again preventing addition or loss of molecule's at that end. Tropomodulin is typically found in cells that require extremely stable actin filaments, such as those in muscle and red blood cells.[120]
These actin binding proteins are typically regulated by various cellular signals to control actin assembly dynamics in different cellular locations. Formins, for example, are typically folded in an inactive conformation until they're activated by the binding of the small GTPase Rho.[117] Actin branching at the cell membrane is important for cell movement, and so the plasma membrane lipid PIP2 activates the nucleation promoting factor WASp and inhibits CapZ.[122] WASp is also activated by the small GTPase Cdc42, while another nucleation promoting factor WAVE is activated by the GTPase Rac1.[123]
Genetics
Although most yeasts have only a single actin gene, higher eukaryotes, in general, express several isoforms of actin encoded by a family of related genes. Mammals have at least six actin isoforms coded by separate genes,[124] which are divided into three classes – alpha, beta, and gamma – according to their isoelectric points. In general, alpha actins are found in muscle (α-skeletal, α-aortic smooth, α-cardiac), whereas beta and gamma isoforms are prominent in non-muscle cells (β-cytoplasmic, γ1-cytoplasmic, γ2-enteric smooth). Although the amino acid sequences and in vitro properties of the isoforms are highly similar, these isoforms cannot completely substitute for one another in vivo.[125] Plants contains more than 60 actin genes and pseudogenes.[84]
The typical actin gene has an approximately 100-nucleotide 5' UTR, a 1200-nucleotide translated region, and a 200-nucleotide 3' UTR. The majority of actin genes are interrupted by introns, with up to six introns in any of 19 well-characterised locations. The high conservation of the family makes actin the favoured model for studies comparing the introns-early and introns-late models of intron evolution.
Evolution
Actin and closely related proteins are present in all organisms, suggesting the common ancestor of all life on Earth had actin.[126] Actin is one of the most conserved proteins throughout the evolution of eukaryotes. The sequences of actin proteins from animals and amoebae are 80% identical despite being separated by approximately one billion years of evolution.[84] Many unicellular eukaryotes have a single actin gene, while multicellular eukaryotes often have several closely related genes that serve specialized functions. Humans have six; plants have 10 or more.[126] In addition to actin, eukaryotes have a large family of actin-related proteins, or "Arps", that share a common ancestor with actin and are called Arp1–Arp11, with Arp1 the most closely related to actin, and Arp11 the least.[126]
Bacteria encode three types of actin: MreB influences cell shape, FtsA cell division, and ParM separation of large plasmids.[126] Some archaea have a bacteria-like MreB gene, while others have an actin gene that more closely resembles eukaryote actin.[126]
The eukaryotic cytoskeleton of organisms among all taxonomic groups have similar components to actin and tubulin. For example, the protein that is coded by the ACTG2 gene in humans is completely equivalent to the homologues present in rats and mice, even though at a nucleotide level the similarity decreases to 92%.[127] However, there are major differences with the equivalents in prokaryotes (FtsZ and MreB), where the similarity between nucleotide sequences is between 40 and 50 % among different bacteria and archaea species. Some authors suggest that the ancestral protein that gave rise to the model eukaryotic actin resembles the proteins present in modern bacterial cytoskeletons.[4][128]
Some authors point out that the behaviour of actin, tubulin, and histone, a protein involved in the stabilization and regulation of DNA, are similar in their ability to bind nucleotides and in their ability of take advantage of Brownian motion. It has also been suggested that they all have a common ancestor.[129] Therefore, evolutionary processes resulted in the diversification of ancestral proteins into the varieties present today, conserving, among others, actins as efficient molecules that were able to tackle essential ancestral biological processes, such as endocytosis.[130]
The Arp2/3 complex is widely found in all eukaryotic organisms.[131]
Equivalents in prokaryotes
The bacterial cytoskeleton contains proteins that are highly similar to actin monomers and polymers. The bacterial protein MreB polymerizes into thin non-helical filaments and occasionally into helical structures similar to F-actin.[93][132] Furthermore, its crystalline structure is very similar to that of G-actin (in terms of its three-dimensional conformation), there are even similarities between the MreB protofilaments and F-actin. The bacterial cytoskeleton also contains the FtsZ proteins, which are similar to tubulin.[133]
Bacteria therefore possess a cytoskeleton with homologous elements to actin (for example, MreB, AlfA, ParM, FtsA, and MamK), even though the amino acid sequence of these proteins diverges from that present in animal cells. However, such proteins have a high degree of structural similarity to eukaryotic actin. The highly dynamic microfilaments formed by the aggregation of MreB and ParM are essential to cell viability and they are involved in cell morphogenesis, chromosome segregation, and cell polarity. ParM is an actin homologue that is coded in a plasmid and it is involved in the regulation of plasmid DNA.[4][134] ParMs from different bacterial plasmids can form astonishingly diverse helical structures comprising two[135][136] or four[137] strands to maintain faithful plasmid inheritance.
In archaea the homologue Ta0583 is even more similar to the eukaryotic actins.[138]
Molecular pathology
The majority of mammals possess six different actin genes. Of these, two code for the cytoskeleton (ACTB and ACTG1) while the other four are involved in skeletal striated muscle (ACTA1), smooth muscle tissue (ACTA2), intestinal muscles (ACTG2) and cardiac muscle (ACTC1). The actin in the cytoskeleton is involved in the pathogenic mechanisms of many infectious agents, including HIV. The vast majority of the mutations that affect actin are point mutations that have a dominant effect, with the exception of six mutations involved in nemaline myopathy. This is because in many cases the mutant of the actin monomer acts as a “cap” by preventing the elongation of F-actin.[87]
Pathology associated with ACTA1
ACTA1 is the gene that codes for the α-isoform of actin that is predominant in human skeletal striated muscles, although it is also expressed in heart muscle and in the thyroid gland.[139] Its DNA sequence consists of seven exons that produce five known transcripts.[140] The majority of these consist of point mutations causing substitution of amino acids. The mutations are in many cases associated with a phenotype that determines the severity and the course of the affliction.[87][140]

The mutation alters the structure and function of skeletal muscles producing one of three forms of myopathy: type 3 nemaline myopathy, congenital myopathy with an excess of thin myofilaments (CM) and congenital myopathy with fibre type disproportion (CMFTD). Mutations have also been found that produce core myopathies.[142] Although their phenotypes are similar, in addition to typical nemaline myopathy some specialists distinguish another type of myopathy called actinic nemaline myopathy. In the former, clumps of actin form instead of the typical rods. It is important to state that a patient can show more than one of these phenotypes in a biopsy.[143] The most common symptoms consist of a typical facial morphology (myopathic facies), muscular weakness, a delay in motor development and respiratory difficulties. The course of the illness, its gravity, and the age at which it appears are all variable and overlapping forms of myopathy are also found. A symptom of nemaline myopathy is that "nemaline rods" appear in differing places in type 1 muscle fibres. These rods are non-pathognomonic structures that have a similar composition to the Z disks found in the sarcomere.[144]
The pathogenesis of this myopathy is very varied. Many mutations occur in the region of actin's indentation near to its nucleotide binding sites, while others occur in Domain 2, or in the areas where interaction occurs with associated proteins. This goes some way to explain the great variety of clumps that form in these cases, such as Nemaline or Intranuclear Bodies or Zebra Bodies.[87] Changes in actin's folding occur in nemaline myopathy as well as changes in its aggregation and there are also changes in the expression of other associated proteins. In some variants where intranuclear bodies are found the changes in the folding masks the nucleus's protein exportation signal so that the accumulation of actin's mutated form occurs in the cell nucleus.[145] On the other hand, it appears that mutations to ACTA1 that give rise to a CFTDM have a greater effect on sarcomeric function than on its structure.[146] Recent investigations have tried to understand this apparent paradox, which suggests there is no clear correlation between the number of rods and muscular weakness. It appears that some mutations are able to induce a greater apoptosis rate in type II muscular fibres.[101]

In smooth muscle
There are two isoforms that code for actins in the smooth muscle tissue:
ACTG2 codes for the largest actin isoform, which has nine exons, one of which, the one located at the 5' end, is not translated.[127] It is a γ-actin that is expressed in the enteric smooth muscle. No mutations to this gene have been found that correspond to pathologies, although microarrays have shown that this protein is more often expressed in cases that are resistant to chemotherapy using cisplatin.[147]
ACTA2 codes for an α-actin located in the smooth muscle, and also in vascular smooth muscle. It has been noted that the MYH11 mutation could be responsible for at least 14% of hereditary thoracic aortic aneurisms particularly Type 6. This is because the mutated variant produces an incorrect filamentary assembly and a reduced capacity for vascular smooth muscle contraction. Degradation of the aortic media has been recorded in these individuals, with areas of disorganization and hyperplasia as well as stenosis of the aorta's vasa vasorum.[148] The number of afflictions that the gene is implicated in is increasing. It has been related to Moyamoya disease and it seems likely that certain mutations in heterozygosis could confer a predisposition to many vascular pathologies, such as thoracic aortic aneurysm and ischaemic heart disease.[149] The α-actin found in smooth muscles is also an interesting marker for evaluating the progress of liver cirrhosis.[150]
In heart muscle
The ACTC1 gene codes for the α-actin isoform present in heart muscle. It was first sequenced by Hamada and co-workers in 1982, when it was found that it is interrupted by five introns.[151] It was the first of the six genes where alleles were found that were implicated in pathological processes.[152]
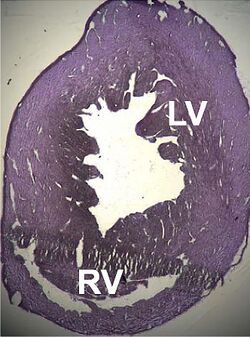
A number of structural disorders associated with point mutations of this gene have been described that cause malfunctioning of the heart, such as Type 1R dilated cardiomyopathy and Type 11 hypertrophic cardiomyopathy. Certain defects of the atrial septum have been described recently that could also be related to these mutations.[154][155]
Two cases of dilated cardiomyopathy have been studied involving a substitution of highly conserved amino acids belonging to the protein domains that bind and intersperse with the Z discs. This has led to the theory that the dilation is produced by a defect in the transmission of contractile force in the myocytes.[156][152]
The mutations in ACTC1 are responsible for at least 5% of hypertrophic cardiomyopathies.[157] The existence of a number of point mutations have also been found:[158]
- Mutation E101K: changes of net charge and formation of a weak electrostatic link in the actomyosin-binding site.
- P166A: interaction zone between actin monomers.
- A333P: actin-myosin interaction zone.
Pathogenesis appears to involve a compensatory mechanism: the mutated proteins act like toxins with a dominant effect, decreasing the heart's ability to contract causing abnormal mechanical behaviour such that the hypertrophy, that is usually delayed, is a consequence of the cardiac muscle's normal response to stress.[159]
Recent studies have discovered ACTC1 mutations that are implicated in two other pathological processes: Infantile idiopathic restrictive cardiomyopathy,[160] and noncompaction of the left ventricular myocardium.[161]
In cytoplasmatic actins
ACTB is a highly complex locus. A number of pseudogenes exist that are distributed throughout the genome, and its sequence contains six exons that can give rise to up to 21 different transcriptions by alternative splicing, which are known as the β-actins. Consistent with this complexity, its products are also found in a number of locations and they form part of a wide variety of processes (cytoskeleton, NuA4 histone-acyltransferase complex, cell nucleus) and in addition they are associated with the mechanisms of a great number of pathological processes (carcinomas, juvenile dystonia, infection mechanisms, nervous system malformations and tumour invasion, among others).[162] A new form of actin has been discovered, kappa actin, which appears to substitute for β-actin in processes relating to tumours.[163]

Three pathological processes have so far been discovered that are caused by a direct alteration in gene sequence:
- Hemangiopericytoma with t(7;12)(p22;q13)-translocations is a rare affliction, in which a translocational mutation causes the fusion of the ACTB gene over GLI1 in Chromosome 12.[165]
- Juvenile onset dystonia is a rare degenerative disease that affects the central nervous system; in particular, it affects areas of the neocortex and thalamus, where rod-like eosinophilic inclusions are formed. The affected individuals represent a phenotype with deformities on the median line, sensory hearing loss and dystonia. It is caused by a point mutation in which the amino acid tryptophan replaces arginine in position 183. This alters actin's interaction with the ADF/cofilin system, which regulates the dynamics of nerve cell cytoskeleton formation.[166]
- A dominant point mutation has also been discovered that causes neutrophil granulocyte dysfunction and recurring infections. It appears that the mutation modifies the domain responsible for binding between profilin and other regulatory proteins. Actin's affinity for profilin is greatly reduced in this allele.[167]
The ACTG1 locus codes for the cytosolic γ-actin protein that is responsible for the formation of cytoskeletal microfilaments. It contains six exons, giving rise to 22 different mRNAs, which produce four complete isoforms whose form of expression is probably dependent on the type of tissue they are found in. It also has two different DNA promoters.[168] It has been noted that the sequences translated from this locus and from that of β-actin are very similar to the predicted ones, suggesting a common ancestral sequence that suffered duplication and genetic conversion.[169]
In terms of pathology, it has been associated with processes such as amyloidosis, retinitis pigmentosa, infection mechanisms, kidney diseases, and various types of congenital hearing loss.[168]
Six autosomal-dominant point mutations in the sequence have been found to cause various types of hearing loss, particularly sensorineural hearing loss linked to the DFNA 20/26 locus. It seems that they affect the stereocilia of the ciliated cells present in the inner ear's Organ of Corti. β-actin is the most abundant protein found in human tissue, but it is not very abundant in ciliated cells, which explains the location of the pathology. On the other hand, it appears that the majority of these mutations affect the areas involved in linking with other proteins, particularly actomyosin.[87] Some experiments have suggested that the pathological mechanism for this type of hearing loss relates to the F-actin in the mutations being more sensitive to cofilin than normal.[170]
However, although there is no record of any case, it is known that γ-actin is also expressed in skeletal muscles, and although it is present in small quantities, model organisms have shown that its absence can give rise to myopathies.[171]
Other pathological mechanisms
Some infectious agents use actin, especially cytoplasmic actin, in their life cycle. Two basic forms are present in bacteria:
- Listeria monocytogenes, some species of Rickettsia, Shigella flexneri and other intracellular germs escape from phagocytic vacuoles by coating themselves with a capsule of actin filaments. L. monocytogenes and S. flexneri both generate a tail in the form of a "comet tail" that gives them mobility. Each species exhibits small differences in the molecular polymerization mechanism of their "comet tails". Different displacement velocities have been observed, for example, with Listeria and Shigella found to be the fastest.[172] Many experiments have demonstrated this mechanism in vitro. This indicates that the bacteria are not using a myosin-like protein motor, and it appears that their propulsion is acquired from the pressure exerted by the polymerization that takes place near to the microorganism's cell wall. The bacteria have previously been surrounded by ABPs from the host, and as a minimum the covering contains Arp2/3 complex, Ena/VASP proteins, cofilin, a buffering protein and nucleation promoters, such as vinculin complex. Through these movements they form protrusions that reach the neighbouring cells, infecting them as well so that the immune system can only fight the infection through cell immunity. The movement could be caused by the modification of the curve and debranching of the filaments.[173] Other species, such as Mycobacterium marinum and Burkholderia pseudomallei, are also capable of localized polymerization of cellular actin to aid their movement through a mechanism that is centered on the Arp2/3 complex. In addition the vaccine virus Vaccinia also uses elements of the actin cytoskeleton for its dissemination.[174]
- Pseudomonas aeruginosa is able to form a protective biofilm in order to escape a host organism’s defences, especially white blood cells and antibiotics. The biofilm is constructed using DNA and actin filaments from the host organism.[175]
In addition to the previously cited example, actin polymerization is stimulated in the initial steps of the internalization of some viruses, notably HIV, by, for example, inactivating the cofilin complex.[176]
The role that actin plays in the invasion process of cancer cells has still not been determined.[177]
Applications
Actin is used in scientific and technological laboratories as a track for molecular motors such as myosin (either in muscle tissue or outside it) and as a necessary component for cellular functioning. It can also be used as a diagnostic tool, as several of its anomalous variants are related to the appearance of specific pathologies.
- Nanotechnology. Actin-myosin systems act as molecular motors that permit the transport of vesicles and organelles throughout the cytoplasm. It is possible that actin could be applied to nanotechnology as its dynamic ability has been harnessed in a number of experiments including those carried out in acellular systems. The underlying idea is to use the microfilaments as tracks to guide molecular motors that can transport a given load. That is actin could be used to define a circuit along which a load can be transported in a more or less controlled and directed manner. In terms of general applications, it could be used for the directed transport of molecules for deposit in determined locations, which would permit the controlled assembly of nanostructures.[178] These attributes could be applied to laboratory processes such as on lab-on-a-chip, in nanocomponent mechanics and in nanotransformers that convert mechanical energy into electrical energy.[179]

- Actin is used as an internal control in western blots to ascertain that equal amounts of protein have been loaded on each lane of the gel. In the blot example shown on the left side, 75 µg of total protein was loaded in each well. The blot was reacted with anti-β-actin antibody (for other details of the blot see the reference [180])
The use of actin as an internal control is based on the assumption that its expression is practically constant and independent of experimental conditions. By comparing the expression of the gene of interest to that of the actin, it is possible to obtain a relative quantity that can be compared between different experiments,[181] whenever the expression of the latter is constant. It is worth pointing out that actin does not always have the desired stability in its gene expression.[182]
- Health. Some alleles of actin cause diseases; for this reason techniques for their detection have been developed. In addition, actin can be used as an indirect marker in surgical pathology: it is possible to use variations in the pattern of its distribution in tissue as a marker of invasion in neoplasia, vasculitis, and other conditions.[183] Further, due to actin's close association with the apparatus of muscular contraction its levels in skeletal muscle diminishes when these tissues atrophy, it can therefore be used as a marker of this physiological process.[184]
- Food technology. It is possible to determine the quality of certain processed foods, such as sausages, by quantifying the amount of actin present in the constituent meat. Traditionally, a method has been used that is based on the detection of 3-methylhistidine in hydrolyzed samples of these products, as this compound is present in actin and F-myosin's heavy chain (both are major components of muscle). The generation of this compound in flesh derives from the methylation of histidine residues present in both proteins.[185][186]
History

Actin was first observed experimentally in 1887 by W.D. Halliburton, who extracted a protein from muscle that 'coagulated' preparations of myosin that he called "myosin-ferment".[187] However, Halliburton was unable to further refine his findings, and the discovery of actin is credited instead to Brunó Ferenc Straub, a young biochemist working in Albert Szent-Györgyi's laboratory at the Institute of Medical Chemistry at the University of Szeged, Hungary.
Following up on the discovery of Ilona Banga & Szent-Györgyi in 1941 that the coagulation only occurs in some myosin extractions and was reversed upon the addition of ATP,[188] Straub identified and purified actin from those myosin preparations that did coagulate. Building on Banga's original extraction method, he developed a novel technique for extracting muscle protein that allowed him to isolate substantial amounts of relatively pure actin, published in 1942.[189] Straub's method is essentially the same as that used in laboratories today. Since Straub's protein was necessary to activate the coagulation of myosin, it was dubbed actin.[188][190] Realizing that Banga's coagulating myosin preparations contained actin as well, Szent-Györgyi called the mixture of both proteins actomyosin.[191]
The hostilities of World War II meant Szent-Gyorgyi was unable to publish his lab's work in Western scientific journals. Actin therefore only became well known in the West in 1945, when their paper was published as a supplement to the Acta Physiologica Scandinavica.[192] Straub continued to work on actin, and in 1950 reported that actin contains bound ATP[193] and that, during polymerization of the protein into microfilaments, the nucleotide is hydrolyzed to ADP and inorganic phosphate (which remain bound to the microfilament). Straub suggested that the transformation of ATP-bound actin to ADP-bound actin played a role in muscular contraction. In fact, this is true only in smooth muscle, and was not supported through experimentation until 2001.[193][194]
The amino acid sequencing of actin was completed by M. Elzinga and co-workers in 1973.[85] The crystal structure of G-actin was solved in 1990 by Kabsch and colleagues.[88] In the same year, a model for F-actin was proposed by Holmes and colleagues following experiments using co-crystallization with different proteins.[90] The procedure of co-crystallization with different proteins was used repeatedly during the following years, until in 2001 the isolated protein was crystallized along with ADP. However, there is still no high-resolution X-ray structure of F-actin. The crystallization of G-actin was possible due to the use of a rhodamine conjugate that impedes polymerization by blocking the amino acid cys-374.[1] Christine Oriol-Audit died in the same year that actin was first crystallized but she was the researcher that in 1977 first crystallized actin in the absence of Actin Binding Proteins (ABPs). However, the resulting crystals were too small for the available technology of the time.[195]
Although no high-resolution model of actin's filamentous form currently exists, in 2008 Sawaya's team were able to produce a more exact model of its structure based on multiple crystals of actin dimers that bind in different places.[196] This model has subsequently been further refined by Sawaya and Lorenz. Other approaches such as the use of cryo-electron microscopy and synchrotron radiation have recently allowed increasing resolution and better understanding of the nature of the interactions and conformational changes implicated in the formation of actin filaments.[197][93][96]
Research
Chemical inhibitors

A number of natural toxins that interfere with actin's dynamics are widely used in research to study actin's role in biology. Latrunculin – a toxin produced by sponges – binds to G-actin preventing it from joining microfilaments.[198] Cytochalasin D – produced by certain fungi – serves as a capping factor, binding to the (+) end of a filament and preventing further addition of actin molecules.[198] In contrast, the sponge toxin jasplakinolide promotes the nucleation of new actin filaments by binding and stabilzing pairs of actin molecules.[199] Phalloidin – from the "death cap" mushroom Amanita phalloides – binds to adjacent actin molecules within the F-actin filament, stabilizing the filament and preventing its depolymerization.[199]
Phalloidin is often labelled with fluorescent dyes to visualize actin filaments by fluorescence microscopy.[199]
See also
- Actin remodeling — effect on cell structure and shape
- Active matter
- Filopodia
- Intermediate filament
- Lamellipodium
- Motor protein — converts chemical energy into mechanical work
- Neuron
- Phallotoxin
References
- ↑ 1.0 1.1 1.2 1.3 1.4 PDB: 1J6Z; "The crystal structure of uncomplexed actin in the ADP state". Science 293 (5530): 708–711. Jul 2001. doi:10.1126/science.1059700. PMID 11474115.
- ↑ "Mediation, modulation, and consequences of membrane-cytoskeleton interactions". Annual Review of Biophysics 37 (1): 65–95. 2008. doi:10.1146/annurev.biophys.37.032807.125912. PMID 18573073.
- ↑ "Cytoskeletal tropomyosins: choreographers of actin filament functional diversity". Journal of Muscle Research and Cell Motility 34 (3–4): 261–274. Aug 2013. doi:10.1007/s10974-013-9355-8. PMID 23904035.
- ↑ 4.0 4.1 4.2 4.3 "The evolution of compositionally and functionally distinct actin filaments". Journal of Cell Science 128 (11): 2009–2019. Jun 2015. doi:10.1242/jcs.165563. PMID 25788699.
- ↑ "In search of the primordial actin filament". Proceedings of the National Academy of Sciences of the United States of America 112 (30): 9150–9151. Jul 2015. doi:10.1073/pnas.1511568112. PMID 26178194.
- ↑ 6.0 6.1 "Chapter 16: The cytoskeleton". Molecular biology of the cell. New York: Garland Science. 2002. pp. 907–982. ISBN 978-0-8153-3218-3. https://archive.org/details/molecularbiolog000wils.
- ↑ 7.0 7.1 7.2 Lodish et al. 2016, p. 778.
- ↑ "Emergent complexity of the cytoskeleton: from single filaments to tissue". Advances in Physics 62 (1): 1–112. January 2013. doi:10.1080/00018732.2013.771509. PMID 24748680. Bibcode: 2013AdPhy..62....1H.
- ↑ 9.0 9.1 9.2 "Actin and myosin as transcription factors". Current Opinion in Genetics & Development 16 (2): 191–196. Apr 2006. doi:10.1016/j.gde.2006.02.001. PMID 16495046.
- ↑ Eckert animal physiology: mechanisms and adaptations. New York: W.H. Freeman and CO. 2002. ISBN 978-0-7167-3863-3. https://archive.org/details/eckertanimalphys00rand.
- ↑ 11.0 11.1 (in es) Citología e histología vegetal y animal. McGraw-Hill Interamericana de España, S.A.U.. 2002. ISBN 978-84-486-0436-3.
- ↑ "Actin, spectrin, and associated proteins form a periodic cytoskeletal structure in axons". Science 339 (6118): 452–456. Jan 2013. doi:10.1126/science.1232251. PMID 23239625. Bibcode: 2013Sci...339..452X.
- ↑ 13.0 13.1 "The yeast actin cytoskeleton: from cellular function to biochemical mechanism". Microbiology and Molecular Biology Reviews 70 (3): 605–645. Sep 2006. doi:10.1128/MMBR.00013-06. PMID 16959963.
- ↑ "Isovariant dynamics expand and buffer the responses of complex systems: the diverse plant actin gene family". The Plant Cell 11 (6): 995–1006. Jun 1999. doi:10.1105/tpc.11.6.995. PMID 10368172.
- ↑ PDB 1unc; "Solution structures of the C-terminal headpiece subdomains of human villin and advillin, evaluation of headpiece F-actin-binding requirements". Protein Science 13 (5): 1276–1287. May 2004. doi:10.1110/ps.03518104. PMID 15096633.
- ↑ 16.0 16.1 "Actin microfilament dynamics and actin side-binding proteins in plants". Current Opinion in Plant Biology 10 (6): 549–556. Dec 2007. doi:10.1016/j.pbi.2007.08.012. PMID 17936064.
- ↑ "AtFim1 is an actin filament crosslinking protein from Arabidopsis thaliana". The Plant Journal 24 (5): 625–636. Dec 2000. doi:10.1046/j.1365-313x.2000.00907.x. PMID 11123801.
- ↑ 18.0 18.1 "Diffusible and bound actin nuclei of Xenopus laevis oocytes". Cell 12 (4): 883–891. Dec 1977. doi:10.1016/0092-8674(77)90152-0. PMID 563771.
- ↑ Cell and molecular biology of nuclear actin. International Review of Cell and Molecular Biology. 273. 2009-01-01. pp. 219–263. doi:10.1016/S1937-6448(08)01806-6. ISBN 9780123748041.
- ↑ "Emerging properties and functions of actin and actin filaments inside the nucleus". Cold Spring Harbor Perspectives in Biology 13 (3): a040121. Feb 2021. doi:10.1101/cshperspect.a040121. PMID 33288541.
- ↑ 21.0 21.1 "A selective block of nuclear actin export stabilizes the giant nuclei of Xenopus oocytes". Nature Cell Biology 8 (3): 257–263. Mar 2006. doi:10.1038/ncb1357. PMID 16489345.
- ↑ "Active maintenance of nuclear actin by importin 9 supports transcription". Proceedings of the National Academy of Sciences of the United States of America 109 (9): E544–552. Feb 2012. doi:10.1073/pnas.1118880109. PMID 22323606.
- ↑ "Nuclear export of actin: a novel mechanism regulating the subcellular localization of a major cytoskeletal protein". The EMBO Journal 17 (6): 1635–1641. Mar 1998. doi:10.1093/emboj/17.6.1635. PMID 9501085.
- ↑ "Exportin 6: a novel nuclear export receptor that is specific for profilin.actin complexes". The EMBO Journal 22 (21): 5928–5940. Nov 2003. doi:10.1093/emboj/cdg565. PMID 14592989.
- ↑ "SUMOylation of nuclear actin". The Journal of Cell Biology 186 (2): 193–200. Jul 2009. doi:10.1083/jcb.200905016. PMID 19635839.
- ↑ "Nucleoplasmic beta-actin exists in a dynamic equilibrium between low-mobility polymeric species and rapidly diffusing populations". The Journal of Cell Biology 172 (4): 541–552. Feb 2006. doi:10.1083/jcb.200507101. PMID 16476775.
- ↑ "Tracking down the different forms of nuclear actin". Trends in Cell Biology 16 (8): 391–396. Aug 2006. doi:10.1016/j.tcb.2006.06.006. PMID 16828286.
- ↑ 28.0 28.1 28.2 28.3 "β- and γ-Actins in the nucleus of human melanoma A375 cells". Histochemistry and Cell Biology 144 (5): 417–428. Nov 2015. doi:10.1007/s00418-015-1349-8. PMID 26239425.
- ↑ "Actin in the nucleus: what form and what for?". Journal of Structural Biology 140 (1–3): 3–9. 2002-12-01. doi:10.1016/s1047-8477(02)00528-2. PMID 12490148.
- ↑ "Cofilin, actin and their complex observed in vivo using fluorescence resonance energy transfer". Biophysical Journal 89 (3): 1902–1908. Sep 2005. doi:10.1529/biophysj.105.062083. PMID 15994898. Bibcode: 2005BpJ....89.1902C.
- ↑ "Nuclear actin: A key player in extracellular matrix-nucleus communication". Communicative & Integrative Biology 4 (5): 511–512. Sep 2011. doi:10.4161/cib.16256. PMID 22046450.
- ↑ 32.0 32.1 "Rapid and phosphoinositol-dependent binding of the SWI/SNF-like BAF complex to chromatin after T lymphocyte receptor signaling". Cell 95 (5): 625–636. Nov 1998. doi:10.1016/s0092-8674(00)81633-5. PMID 9845365.
- ↑ 33.0 33.1 "Actin is part of pre-initiation complexes and is necessary for transcription by RNA polymerase II". Nature Cell Biology 6 (11): 1094–1101. Nov 2004. doi:10.1038/ncb1182. PMID 15502823.
- ↑ 34.0 34.1 "A role for beta-actin in RNA polymerase III transcription". Genes & Development 18 (24): 3010–3015. Dec 2004. doi:10.1101/gad.1250804. PMID 15574586.
- ↑ 35.0 35.1 "Nuclear actin and myosin I are required for RNA polymerase I transcription". Nature Cell Biology 6 (12): 1165–1172. Dec 2004. doi:10.1038/ncb1190. PMID 15558034.
- ↑ "New roles for lamins, nuclear envelope proteins and actin in the nucleus". Advances in Enzyme Regulation 44: 155–172. 2004-01-01. doi:10.1016/j.advenzreg.2003.11.005. PMID 15581488.
- ↑ "Cells lacking β-actin are genetically reprogrammed and maintain conditional migratory capacity". Molecular & Cellular Proteomics 11 (8): 255–271. Aug 2012. doi:10.1074/mcp.M111.015099. PMID 22448045.
- ↑ "Emerin caps the pointed end of actin filaments: evidence for an actin cortical network at the nuclear inner membrane". PLOS Biology 2 (9): E231. Sep 2004. doi:10.1371/journal.pbio.0020231. PMID 15328537.
- ↑ "Emery–Dreifuss muscular dystrophy". Muscular Dystrophies. Handbook of Clinical Neurology. 101. 2011-01-01. pp. 155–166. doi:10.1016/B978-0-08-045031-5.00012-8. ISBN 9780080450315.
- ↑ "Chromatin remodelling and actin organisation". FEBS Letters 582 (14): 2041–2050. Jun 2008. doi:10.1016/j.febslet.2008.04.032. PMID 18442483.
- ↑ "The growing pre-mRNA recruits actin and chromatin-modifying factors to transcriptionally active genes". Genes & Development 19 (16): 1871–1884. Aug 2005. doi:10.1101/gad.339405. PMID 16103215.
- ↑ 42.0 42.1 "Molecular functions of nuclear actin in transcription". The Journal of Cell Biology 172 (7): 967–971. Mar 2006. doi:10.1083/jcb.200512083. PMID 16549500.
- ↑ "Nuclear architecture and gene regulation". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research 1783 (11): 2174–2184. Nov 2008. doi:10.1016/j.bbamcr.2008.07.018. PMID 18718493.
- ↑ "Actin on DNA-an ancient and dynamic relationship". Cytoskeleton 67 (8): 487–495. Aug 2010. doi:10.1002/cm.20464. PMID 20593452.
- ↑ "Nuclear actin and actin-related proteins in chromatin remodeling". Annual Review of Biochemistry 71: 755–781. 2002-01-01. doi:10.1146/annurev.biochem.71.110601.135507. PMID 12045110.
- ↑ "Nuclear actin and actin-binding proteins in the regulation of transcription and gene expression". The FEBS Journal 276 (10): 2669–2685. May 2009. doi:10.1111/j.1742-4658.2009.06986.x. PMID 19459931.
- ↑ "Induction of HoxB transcription by retinoic acid requires actin polymerization". Molecular Biology of the Cell 20 (15): 3543–3551. Aug 2009. doi:10.1091/mbc.E09-02-0114. PMID 19477923.
- ↑ "Nuclear translocation of beta-actin is involved in transcriptional regulation during macrophage differentiation of HL-60 cells". Molecular Biology of the Cell 21 (5): 811–820. Mar 2010. doi:10.1091/mbc.E09-06-0534. PMID 20053683.
- ↑ 49.0 49.1 "Nuclear actin polymerization is required for transcriptional reprogramming of Oct4 by oocytes". Genes & Development 25 (9): 946–958. May 2011. doi:10.1101/gad.615211. PMID 21536734.
- ↑ "Coronin 2A mediates actin-dependent de-repression of inflammatory response genes". Nature 470 (7334): 414–418. Feb 2011. doi:10.1038/nature09703. PMID 21331046. Bibcode: 2011Natur.470..414H.
- ↑ "Nuclear actin and transcriptional activation". Communicative & Integrative Biology 4 (5): 582–583. Sep 2011. doi:10.4161/cib.16491. PMID 22046469.
- ↑ "Long-range directional movement of an interphase chromosome site". Current Biology 16 (8): 825–831. Apr 2006. doi:10.1016/j.cub.2006.03.059. PMID 16631592.
- ↑ "Nuclear myosin I is necessary for the formation of the first phosphodiester bond during transcription initiation by RNA polymerase II". Journal of Cellular Biochemistry 99 (4): 1001–1009. Nov 2006. doi:10.1002/jcb.21035. PMID 16960872.
- ↑ "Linking actin dynamics and gene transcription to drive cellular motile functions". Nature Reviews Molecular Cell Biology 11 (5): 353–365. May 2010. doi:10.1038/nrm2890. PMID 20414257.
- ↑ "Actin dynamics control SRF activity by regulation of its coactivator MAL". Cell 113 (3): 329–342. May 2003. doi:10.1016/s0092-8674(03)00278-2. PMID 12732141.
- ↑ "Nuclear actin dynamics--from form to function". FEBS Letters 582 (14): 2033–2040. Jun 2008. doi:10.1016/j.febslet.2008.04.010. PMID 18423404.
- ↑ "Actin-mediated gene expression in neurons: the MRTF-SRF connection". Biological Chemistry 391 (6): 591–597. Jun 2010. doi:10.1515/BC.2010.061. PMID 20370316.
- ↑ "T cell receptor-triggered nuclear actin network formation drives CD4+ T cell effector functions". Science Immunology 4 (31): eaav1987. Jan 2019. doi:10.1126/sciimmunol.aav1987. PMID 30610013.
- ↑ 59.0 59.1 59.2 59.3 Lodish et al. 2016, p. 811–812.
- ↑ Lodish et al. 2016, p. 796.
- ↑ 61.0 61.1 61.2 61.3 61.4 Lodish et al. 2016, p. 803–805.
- ↑ Lodish et al. 2016, p. 805–806.
- ↑ 63.0 63.1 Lodish et al. 2016, p. 807.
- ↑ "Alpha-actinin localization in the cleavage furrow during cytokinesis". The Journal of Cell Biology 79 (1): 268–275. Oct 1978. doi:10.1083/jcb.79.1.268. PMID 359574.
- ↑ "Actin dynamics in the contractile ring during cytokinesis in fission yeast". Nature 419 (6902): 82–86. Sep 2002. doi:10.1038/nature00999. PMID 12214236. Bibcode: 2002Natur.419...82P.
- ↑ Lodish et al. 2016, p. 809.
- ↑ "Actin cleavage by CPP-32/apopain during the development of apoptosis". Oncogene 14 (9): 1007–1012. Mar 1997. doi:10.1038/sj.onc.1200919. PMID 9070648.
- ↑ "Calpain and caspase: can you tell the difference?". Trends in Neurosciences 23 (1): 20–26. Jan 2000. doi:10.1016/S0166-2236(99)01479-4. PMID 10631785.
- ↑ "Calpain inhibitors, but not caspase inhibitors, prevent actin proteolysis and DNA fragmentation during apoptosis". Journal of Cell Science 111 (Pt 6): 713–722. Mar 1998. doi:10.1242/jcs.111.6.713. PMID 9472000.
- ↑ "SAPK2/p38-dependent F-actin reorganization regulates early membrane blebbing during stress-induced apoptosis". The Journal of Cell Biology 143 (5): 1361–1373. Nov 1998. doi:10.1083/jcb.143.5.1361. PMID 9832563.
- ↑ "Quantitative analysis of cadherin-catenin-actin reorganization during development of cell-cell adhesion". The Journal of Cell Biology 135 (6 Pt 2): 1899–1911. Dec 1996. doi:10.1083/jcb.135.6.1899. PMID 8991100.
- ↑ "Redundancy in the microfilament system: abnormal development of Dictyostelium cells lacking two F-actin cross-linking proteins". Cell 68 (1): 53–62. Jan 1992. doi:10.1016/0092-8674(92)90205-Q. PMID 1732064.
- ↑ "Actin plays a role in both changes in cell shape and gene-expression associated with Schwann cell myelination". The Journal of Neuroscience 17 (1): 241–250. Jan 1997. doi:10.1523/JNEUROSCI.17-01-00241.1997. PMID 8987752.
- ↑ "Role of actin cytoskeletal dynamics in activation of the cyclic AMP pathway and HWP1 gene expression in Candida albicans". Eukaryotic Cell 6 (10): 1824–1840. Oct 2007. doi:10.1128/EC.00188-07. PMID 17715368.
- ↑ "Novel actin-like proteins T-ACTIN 1 and T-ACTIN 2 are differentially expressed in the cytoplasm and nucleus of mouse haploid germ cells". Biology of Reproduction 69 (2): 475–482. Aug 2003. doi:10.1095/biolreprod.103.015867. PMID 12672658.
- ↑ "Epigenetic effects on yeast transcription caused by mutations in an actin-related protein present in the nucleus". Genes & Development 10 (5): 604–619. Mar 1996. doi:10.1101/gad.10.5.604. PMID 8598290.
- ↑ "Dynamic length regulation of sensory stereocilia". Seminars in Cell & Developmental Biology 19 (6): 502–510. Dec 2008. doi:10.1016/j.semcdb.2008.07.006. PMID 18692583.
- ↑ "An actin molecular treadmill and myosins maintain stereocilia functional architecture and self-renewal". The Journal of Cell Biology 164 (6): 887–897. Mar 2004. doi:10.1083/jcb.200310055. PMID 15024034.
- ↑ "Polarity reveals intrinsic cell chirality". Proceedings of the National Academy of Sciences of the United States of America 104 (22): 9296–9300. May 2007. doi:10.1073/pnas.0703153104. PMID 17517645. Bibcode: 2007PNAS..104.9296X.
- ↑ "Autonomous right-screw rotation of growth cone filopodia drives neurite turning". The Journal of Cell Biology 188 (3): 429–441. Feb 2010. doi:10.1083/jcb.200906043. PMID 20123994.
- ↑ "Micropatterned mammalian cells exhibit phenotype-specific left-right asymmetry". Proceedings of the National Academy of Sciences of the United States of America 108 (30): 12295–12300. Jul 2011. doi:10.1073/pnas.1103834108. PMID 21709270. Bibcode: 2011PNAS..10812295W.
- ↑ "Crystal structure of monomeric actin in the ATP state. Structural basis of nucleotide-dependent actin dynamics". The Journal of Biological Chemistry 278 (36): 34172–34180. Sep 2003. doi:10.1074/jbc.M303689200. PMID 12813032.
- ↑ "Actin molecular structure and function". Current Opinion in Cell Biology 5 (1): 41–47. Feb 1993. doi:10.1016/S0955-0674(05)80006-7. PMID 8448029.
- ↑ 84.0 84.1 84.2 84.3 84.4 Lodish et al. 2016, p. 779.
- ↑ 85.0 85.1 85.2 "Complete amino-acid sequence of actin of rabbit skeletal muscle". Proceedings of the National Academy of Sciences of the United States of America 70 (9): 2687–2691. Sep 1973. doi:10.1073/pnas.70.9.2687. PMID 4517681. Bibcode: 1973PNAS...70.2687E.
- ↑ 86.0 86.1 "The primary structure of actin from rabbit skeletal muscle. Completion and analysis of the amino acid sequence". The Journal of Biological Chemistry 250 (15): 5915–5920. Aug 1975. doi:10.1016/S0021-9258(19)41139-3. PMID 1150665.
- ↑ 87.0 87.1 87.2 87.3 87.4 87.5 87.6 Actin-binding Proteins and Disease. Springer. 2008. ISBN 978-0-387-71747-0. https://books.google.com/books?id=Y6xHVnH8iOIC&q=actinopathy&pg=PA23.
- ↑ 88.0 88.1 "Atomic structure of the actin:DNase I complex". Nature 347 (6288): 37–44. Sep 1990. doi:10.1038/347037a0. PMID 2395459. Bibcode: 1990Natur.347...37K.
- ↑ "Crystal structures of expressed non-polymerizable monomeric actin in the ADP and ATP states". The Journal of Biological Chemistry 281 (42): 31909–31919. Oct 2006. doi:10.1074/jbc.M601973200. PMID 16920713.
- ↑ 90.0 90.1 "Atomic model of the actin filament". Nature 347 (6288): 44–49. Sep 1990. doi:10.1038/347044a0. PMID 2395461. Bibcode: 1990Natur.347...44H.
- ↑ 91.0 91.1 91.2 Lodish et al. 2016, p. 780.
- ↑ 92.0 92.1 92.2 "Actin structure and function: what we still do not understand". The Journal of Biological Chemistry 282 (50): 36133–36137. Dec 2007. doi:10.1074/jbc.R700030200. PMID 17965017.
- ↑ 93.0 93.1 93.2 93.3 "The nature of the globular- to fibrous-actin transition". Nature 457 (7228): 441–445. Jan 2009. doi:10.1038/nature07685. PMID 19158791. Bibcode: 2009Natur.457..441O.
- ↑ "The visualization of actin filament polarity in thin sections. Evidence for the uniform polarity of membrane-associated filaments". The Journal of Cell Biology 79 (3): 846–852. Dec 1978. doi:10.1083/jcb.79.3.846. PMID 569662.
- ↑ Histologi. Munksgaard. 1981. p. 105. ISBN 978-87-16-08418-7. https://books.google.com/books?id=-C3MOAAACAAJ.
- ↑ 96.0 96.1 96.2 "Structure of the F-actin-tropomyosin complex". Nature 519 (7541): 114–117. May 2015. doi:10.1038/nature14033. PMID 25470062. Bibcode: 2015Natur.519..114V.
- ↑ Textbook of medical physiology. St. Louis, Mo: Elsevier Saunders. 2006. p. 76. ISBN 978-0-7216-0240-0. https://archive.org/details/textbookmedicalp09guyt.
- ↑ "The Self-Assembly and Dynamic Structure of Cytoskeletal Filaments". Molecular Biology of the Cell (4th ed.). Garland Science. 2002. ISBN 978-0-8153-3218-3. https://www.ncbi.nlm.nih.gov/books/NBK26862/.
- ↑ 99.0 99.1 "Selective contribution of eukaryotic prefoldin subunits to actin and tubulin binding". The Journal of Biological Chemistry 279 (6): 4196–4203. Feb 2004. doi:10.1074/jbc.M306053200. PMID 14634002.
- ↑ 100.0 100.1 "Structure of eukaryotic prefoldin and of its complexes with unfolded actin and the cytosolic chaperonin CCT". The EMBO Journal 21 (23): 6377–6386. Dec 2002. doi:10.1093/emboj/cdf640. PMID 12456645.
- ↑ 101.0 101.1 "alpha-Skeletal muscle actin nemaline myopathy mutants cause cell death in cultured muscle cells". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research 1793 (7): 1259–1271. Jul 2009. doi:10.1016/j.bbamcr.2009.04.004. PMID 19393268.
- ↑ 102.0 102.1 "Activities of the chaperonin containing TCP-1 (CCT): implications for cell cycle progression and cytoskeletal organisation". Cell Stress & Chaperones 14 (1): 23–31. Jan 2009. doi:10.1007/s12192-008-0057-x. PMID 18595008.
- ↑ 103.0 103.1 "PhLP3 modulates CCT-mediated actin and tubulin folding via ternary complexes with substrates". The Journal of Biological Chemistry 281 (11): 7012–7021. Mar 2006. doi:10.1074/jbc.M513235200. PMID 16415341.
- ↑ "Prefoldin-nascent chain complexes in the folding of cytoskeletal proteins". The Journal of Cell Biology 145 (2): 265–277. Apr 1999. doi:10.1083/jcb.145.2.265. PMID 10209023.
- ↑ "The inter-ring arrangement of the cytosolic chaperonin CCT". EMBO Reports 8 (3): 252–257. Mar 2007. doi:10.1038/sj.embor.7400894. PMID 17304242.
- ↑ "Actin interacts with CCT via discrete binding sites: a binding transition-release model for CCT-mediated actin folding". Journal of Molecular Biology 355 (1): 124–138. Jan 2006. doi:10.1016/j.jmb.2005.10.051. PMID 16300788.
- ↑ "Actin polymerization kinetics, cap structure, and fluctuations". Proceedings of the National Academy of Sciences of the United States of America 102 (24): 8543–8548. Jun 2005. doi:10.1073/pnas.0501435102. PMID 15939882. Bibcode: 2005PNAS..102.8543V.
- ↑ "Insights into the cooperative nature of ATP hydrolysis in actin filaments". Biophysical Journal 115 (8): 1589–1602. September 2018. doi:10.1016/j.bpj.2018.08.034. PMID 30249402. Bibcode: 2018BpJ...115.1589K.
- ↑ "Unraveling the mystery of ATP hydrolysis in actin filaments". Journal of the American Chemical Society 136 (37): 13053–13058. September 2014. doi:10.1021/ja507169f. PMID 25181471.
- ↑ "Water molecules in the nucleotide binding cleft of actin: effects on subunit conformation and implications for ATP hydrolysis". Journal of Molecular Biology 413 (1): 279–291. October 2011. doi:10.1016/j.jmb.2011.07.068. PMID 21856312.
- ↑ 111.0 111.1 Lodish et al. 2016, p. 781.
- ↑ 112.0 112.1 Lodish et al. 2016, p. 783.
- ↑ 113.0 113.1 Lodish et al. 2016, p. 784.
- ↑ (in es) Biología celular. Elsevier España. 2002. p. 132. ISBN 978-84-458-1105-4. https://books.google.com/books?id=54vSCCv33pYC.
- ↑ "Actin-binding proteins--a unifying hypothesis". Trends in Biochemical Sciences 29 (11): 572–578. Nov 2004. doi:10.1016/j.tibs.2004.09.004. PMID 15501675.
- ↑ "Crystal structure of Arp2/3 complex". Science 294 (5547): 1679–1684. Nov 2001. doi:10.1126/science.1066333. PMID 11721045. Bibcode: 2001Sci...294.1679R.
- ↑ 117.0 117.1 117.2 117.3 Lodish et al. 2016, p. 786.
- ↑ Lodish et al. 2016, p. 787–788.
- ↑ PDB: 2BTF; "The structure of crystalline profilin-beta-actin". Nature 365 (6449): 810–816. Oct 1993. doi:10.1038/365810a0. PMID 8413665. Bibcode: 1993Natur.365..810S.
- ↑ 120.0 120.1 120.2 120.3 Lodish et al. 2016, p. 785.
- ↑ Lodish et al. 2016, pp. 784–785.
- ↑ Lodish et al. 2016, pp. 785, 788.
- ↑ Lodish et al. 2016, pp. 788–789.
- ↑ "At least six different actins are expressed in a higher mammal: an analysis based on the amino acid sequence of the amino-terminal tryptic peptide". Journal of Molecular Biology 126 (4): 783–802. Dec 1978. doi:10.1016/0022-2836(78)90020-7. PMID 745245.
- ↑ Functional specificity of actin isoforms. International Review of Cytology. 202. 2001. pp. 35–98. doi:10.1016/S0074-7696(01)02003-4. ISBN 9780123646064.
- ↑ 126.0 126.1 126.2 126.3 126.4 Pollard 2016, "Genes, sequence conservation, distribution, and abundance".
- ↑ 127.0 127.1 "Structure, chromosome location, and expression of the human smooth muscle (enteric type) gamma-actin gene: evolution of six human actin genes". Molecular and Cellular Biology 11 (6): 3296–3306. Jun 1991. doi:10.1128/mcb.11.6.3296. PMID 1710027.
- ↑ "Evolution of the cytoskeleton". BioEssays 29 (7): 668–677. Jul 2007. doi:10.1002/bies.20601. PMID 17563102.
- ↑ "Are histones, tubulin, and actin derived from a common ancestral protein?". Protoplasma 233 (1–2): 1–5. 2008. doi:10.1007/s00709-008-0305-z. PMID 18615236.
- ↑ "Actin and endocytosis: mechanisms and phylogeny". Current Opinion in Cell Biology 21 (1): 20–27. Feb 2009. doi:10.1016/j.ceb.2009.01.006. PMID 19186047.
- ↑ "Structure and function of the Arp2/3 complex". Current Opinion in Structural Biology 9 (2): 244–249. Apr 1999. doi:10.1016/S0959-440X(99)80034-7. PMID 10322212.
- ↑ "Filament structure, organization, and dynamics in MreB sheets". The Journal of Biological Chemistry 285 (21): 15858–15865. May 2010. doi:10.1074/jbc.M109.095901. PMID 20223832.
- ↑ "Prokaryotic origin of the actin cytoskeleton". Nature 413 (6851): 39–44. Sep 2001. doi:10.1038/35092500. PMID 11544518. Bibcode: 2001Natur.413...39V.
- ↑ "The bacterial actin-like cytoskeleton". Microbiology and Molecular Biology Reviews 70 (4): 888–909. Dec 2006. doi:10.1128/MMBR.00014-06. PMID 17158703.
- ↑ "Structure and filament dynamics of the pSK41 actin-like ParM protein: implications for plasmid DNA segregation". The Journal of Biological Chemistry 285 (13): 10130–10140. Mar 2010. doi:10.1074/jbc.M109.071613. PMID 20106979.
- ↑ "Polymeric structures and dynamic properties of the bacterial actin AlfA". Journal of Molecular Biology 397 (4): 1031–1041. Apr 2010. doi:10.1016/j.jmb.2010.02.010. PMID 20156449.
- ↑ "Novel actin-like filament structure from Clostridium tetani". The Journal of Biological Chemistry 287 (25): 21121–21129. Jun 2012. doi:10.1074/jbc.M112.341016. PMID 22514279.
- ↑ "An actin homolog of the archaeon Thermoplasma acidophilum that retains the ancient characteristics of eukaryotic actin". Journal of Bacteriology 189 (5): 2039–2045. Mar 2007. doi:10.1128/JB.01454-06. PMID 17189356.
- ↑ "A gene atlas of the mouse and human protein-encoding transcriptomes". Proceedings of the National Academy of Sciences of the United States of America 101 (16): 6062–6067. Apr 2004. doi:10.1073/pnas.0400782101. PMID 15075390. Bibcode: 2004PNAS..101.6062S.
- ↑ 140.0 140.1 "ACTS_HUMAN". UniProt Consortium. https://www.uniprot.org/uniprot/P68133.
- ↑ 141.0 141.1 "Phenotypes of myopathy-related actin mutants in differentiated C2C12 myotubes". BMC Cell Biology 8 (1): 2. 2007. doi:10.1186/1471-2121-8-2. PMID 17227580.
- ↑ "Missense mutations of ACTA1 cause dominant congenital myopathy with cores". Journal of Medical Genetics 41 (11): 842–848. Nov 2004. doi:10.1136/jmg.2004.020271. PMID 15520409.
- ↑ "Muscle disease caused by mutations in the skeletal muscle alpha-actin gene (ACTA1)". Neuromuscular Disorders 13 (7–8): 519–531. Sep 2003. doi:10.1016/S0960-8966(03)00101-9. PMID 12921789.
- ↑ "Nemaline Myopathy". GeneReviews [Internet]. Seattle (WA): University of Washington, Seattle. 2002. https://www.ncbi.nlm.nih.gov/books/NBK1288/.
- ↑ "Evidence for a dominant-negative effect in ACTA1 nemaline myopathy caused by abnormal folding, aggregation and altered polymerization of mutant actin isoforms". Human Molecular Genetics 13 (16): 1727–1743. Aug 2004. doi:10.1093/hmg/ddh185. PMID 15198992.
- ↑ "The pathogenesis of ACTA1-related congenital fiber type disproportion". Annals of Neurology 61 (6): 552–561. Jun 2007. doi:10.1002/ana.21112. PMID 17387733.
- ↑ "Expression microarray analysis reveals genes associated with in vitro resistance to cisplatin in a cell line model". Acta Oncologica 46 (5): 651–658. 2007. doi:10.1080/02841860601156157. PMID 17562441.
- ↑ "Mutations in smooth muscle alpha-actin (ACTA2) lead to thoracic aortic aneurysms and dissections". Nature Genetics 39 (12): 1488–1493. Dec 2007. doi:10.1038/ng.2007.6. PMID 17994018.
- ↑ "Mutations in smooth muscle alpha-actin (ACTA2) cause coronary artery disease, stroke, and Moyamoya disease, along with thoracic aortic disease". American Journal of Human Genetics 84 (5): 617–627. May 2009. doi:10.1016/j.ajhg.2009.04.007. PMID 19409525.
- ↑ "The value of alpha-SMA in the evaluation of hepatic fibrosis severity in hepatitis B infection and cirrhosis development: a histopathological and immunohistochemical study". Histopathology 47 (3): 276–280. Sep 2005. doi:10.1111/j.1365-2559.2005.02226.x. PMID 16115228.
- ↑ "Molecular structure and evolutionary origin of human cardiac muscle actin gene". Proceedings of the National Academy of Sciences of the United States of America 79 (19): 5901–5905. Oct 1982. doi:10.1073/pnas.79.19.5901. PMID 6310553. Bibcode: 1982PNAS...79.5901H.
- ↑ 152.0 152.1 "Actin mutations in dilated cardiomyopathy, a heritable form of heart failure". Science 280 (5364): 750–752. May 1998. doi:10.1126/science.280.5364.750. PMID 9563954. Bibcode: 1998Sci...280..750O.
- ↑ "Pol II-expressed shRNA knocks down Sod2 gene expression and causes phenotypes of the gene knockout in mice". PLOS Genetics 2 (1): e10. Jan 2006. doi:10.1371/journal.pgen.0020010. PMID 16450009.
- ↑ Online Mendelian Inheritance in Man (OMIM) Actin, alpha, cardiac muscle; ACTC1 -102540
- ↑ "Alpha-cardiac actin mutations produce atrial septal defects". Human Molecular Genetics 17 (2): 256–265. Jan 2008. doi:10.1093/hmg/ddm302. PMID 17947298.
- ↑ Bioquimica. Barcelona: Reverté. 2006. ISBN 978-84-291-7208-9. https://books.google.com/books?id=p3DCb9lTLx8C&q=la+miosina+se+une+a+la+actina+residuo&pg=PA1021.
- ↑ Kabaeva Z (11 November 2002). Genetic analysis in hypertrophic cardiomyopathy (Thesis). doi:10.18452/14800.
- ↑ "Inherited and de novo mutations in the cardiac actin gene cause hypertrophic cardiomyopathy". Journal of Molecular and Cellular Cardiology 32 (9): 1687–1694. Sep 2000. doi:10.1006/jmcc.2000.1204. PMID 10966831.
- ↑ "Cardiomiopatía Hipertrófica familiar: Genes, mutaciones y modelos animales. Revisión" (in es). Investigación Clínica 45 (1): 69–100. 2004. http://ve.scielo.org/scielo.php?pid=S0535-51332004000100008&script=sci_abstract.
- ↑ "Idiopathic restrictive cardiomyopathy in children is caused by mutations in cardiac sarcomere protein genes". Heart 94 (11): 1478–1484. Nov 2008. doi:10.1136/hrt.2007.134684. PMID 18467357.
- ↑ "Idiopathic common peroneal nerve palsy--a review of thirteen cases". British Journal of Neurosurgery 5 (1): 7–11. 1991. doi:10.3109/02688699108998440. PMID 1850600.
- ↑ "Gene: ACTB". U.S. National Center for Biotechnology Information (NCBI). https://www.ncbi.nlm.nih.gov/IEB/Research/Acembly/av.cgi?db=human&l=ACTB.
- ↑ "Identification of a novel actin isoform in hepatocellular carcinoma". Hepatology Research 36 (1): 33–39. Sep 2006. doi:10.1016/j.hepres.2006.05.003. PMID 16824795.
- ↑ "Hsp27 and axonal growth in adult sensory neurons in vitro". BMC Neuroscience 6 (1): 24. 2005. doi:10.1186/1471-2202-6-24. PMID 15819993.
- ↑ "Soft tissue tumors: Pericytoma with t(7;12)". University Hospital of Poitiers. http://atlasgeneticsoncology.org/Tumors/Pericytomt0712ID5192.html.
- ↑ "A mutation of beta -actin that alters depolymerization dynamics is associated with autosomal dominant developmental malformations, deafness, and dystonia". American Journal of Human Genetics 78 (6): 947–960. Jun 2006. doi:10.1086/504271. PMID 16685646.
- ↑ "A heterozygous mutation of beta-actin associated with neutrophil dysfunction and recurrent infection". Proceedings of the National Academy of Sciences of the United States of America 96 (15): 8693–8698. Jul 1999. doi:10.1073/pnas.96.15.8693. PMID 10411937. Bibcode: 1999PNAS...96.8693N.
- ↑ 168.0 168.1 "Gene: ACTG1". U.S. National Center for Biotechnology Information (NCBI). https://www.ncbi.nlm.nih.gov/IEB/Research/Acembly/av.cgi?db=human&l=ACTG1.
- ↑ "Nucleotide sequence of the human gamma cytoskeletal actin mRNA: anomalous evolution of vertebrate non-muscle actin genes". Nucleic Acids Research 14 (13): 5275–5294. Jul 1986. doi:10.1093/nar/14.13.5275. PMID 3737401.
- ↑ "Allele-specific effects of human deafness gamma-actin mutations (DFNA20/26) on the actin/cofilin interaction". The Journal of Biological Chemistry 284 (27): 18260–18269. Jul 2009. doi:10.1074/jbc.M109.015818. PMID 19419963.
- ↑ "Cytoplasmic gamma-actin is not required for skeletal muscle development but its absence leads to a progressive myopathy". Developmental Cell 11 (3): 387–397. Sep 2006. doi:10.1016/j.devcel.2006.07.001. PMID 16950128.
- ↑ "A comparative study of the actin-based motilities of the pathogenic bacteria Listeria monocytogenes, Shigella flexneri and Rickettsia conorii". Journal of Cell Science 112 (11): 1697–1708. June 1999. doi:10.1242/jcs.112.11.1697. PMID 10318762.
- ↑ "Listeria comet tails: the actin-based motility machinery at work". Trends in Cell Biology 18 (5): 220–227. May 2008. doi:10.1016/j.tcb.2008.03.001. PMID 18396046.
- ↑ "Actin-based motility of intracellular pathogens". Current Opinion in Microbiology 8 (1): 35–45. Feb 2005. doi:10.1016/j.mib.2004.12.013. PMID 15694855.
- ↑ "Neutrophil enhancement of Pseudomonas aeruginosa biofilm development: human F-actin and DNA as targets for therapy". Journal of Medical Microbiology 58 (Pt 4): 492–502. Apr 2009. doi:10.1099/jmm.0.005728-0. PMID 19273646.
- ↑ "HIV infection of T cells: actin-in and actin-out". Science Signaling 2 (66): pe23. 2009. doi:10.1126/scisignal.266pe23. PMID 19366992. https://zenodo.org/record/1231273.
- ↑ "Actin-based protrusions: promoters or inhibitors of cancer invasion?". Cancer Cell 16 (1): 5–7. Jul 2009. doi:10.1016/j.ccr.2009.06.009. PMID 19573806.
- ↑ "Light-controlled molecular shuttles made from motor proteins carrying cargo on engineered surfaces". Nano Letters 1 (5): 235–239. 2001. doi:10.1021/nl015521e. Bibcode: 2001NanoL...1..235H.
- ↑ "Actin-Based Molecular Motors for Cargo Transportation in Nanotechnology—Potentials and Challenges". IEEE Transactions on Advanced Packaging 28 (4): 547–555. 2005. doi:10.1109/TADVP.2005.858309.
- ↑ "Mapping the sites of localization of epithelial sodium channel (ENaC) and CFTR in segments of the mammalian epididymis". Journal of Molecular Histology 50 (2): 141–154. April 2019. doi:10.1007/s10735-019-09813-3. PMID 30659401.
- ↑ "Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes". Genome Biology 3 (7): RESEARCH0034. Jun 2002. doi:10.1186/gb-2002-3-7-research0034. PMID 12184808.
- ↑ "Beta-actin--an unsuitable internal control for RT-PCR". Molecular and Cellular Probes 15 (5): 307–311. Oct 2001. doi:10.1006/mcpr.2001.0376. PMID 11735303. https://github.com/JeremyMGibson/verbose-garbanzo/raw/master/62818/134000.
- ↑ "Immunohistochemical localization of actin: applications in surgical pathology". The American Journal of Surgical Pathology 5 (1): 91–97. Jan 1981. doi:10.1097/00000478-198101000-00013. PMID 7018275.
- ↑ "Atrophy responses to muscle inactivity. II. Molecular markers of protein deficits". Journal of Applied Physiology 95 (2): 791–802. Aug 2003. doi:10.1152/japplphysiol.01113.2002. PMID 12716877.
- ↑ "Current advances in proteomic analysis and its use for the resolution of poultry meat quality". World's Poultry Science Journal 62 (1): 123–130. 2006. doi:10.1079/WPS200589.
- ↑ "Methods and Instruments in Applied Food Analysis". Handbook of food analysis. 3 (2 ed.). New York, N.Y: Marcel Dekker. 2004. pp. 1741–2226. ISBN 978-0-8247-5039-8.
- ↑ "On Muscle-Plasma". The Journal of Physiology 8 (3–4): 133–202. Aug 1887. doi:10.1113/jphysiol.1887.sp000252. PMID 16991477.
- ↑ 188.0 188.1 "Preparation and properties of myosin A and B.". Studies from the Institute of Medical Chemistry University Szeged. 1941-1942. I: 5–15. 1942. http://actin.aok.pte.hu/archives/index.php.
- ↑ "Actin". Studies from the Institute of Medical Chemistry University Szeged. 1942. II: 3–15. 1942. http://actin.aok.pte.hu/archives/index.php.
- ↑ "The discovery of actin: "to see what everyone else has seen, and to think what nobody has thought"". Journal of Muscle Research and Cell Motility 41 (1): 3–9. March 2020. doi:10.1007/s10974-019-09515-z. PMID 31093826.
- ↑ "Discussion". Studies from the Institute of Medical Chemistry University Szeged. 1941-1942. I: 67–71. 1942. http://actin.aok.pte.hu/archives/index.php.
- ↑ Szent-Gyorgyi A (1945). "Studies on muscle". Acta Physiol Scandinav 9 (Suppl): 25.
- ↑ 193.0 193.1 "Adenosinetriphosphate. The functional group of actin. 1950". Biochimica et Biophysica Acta 1000: 180–195. 1989. doi:10.1016/0006-3002(50)90052-7. PMID 2673365.
- ↑ "Exchange of the actin-bound nucleotide in intact arterial smooth muscle". The Journal of Biological Chemistry 276 (51): 48398–48403. December 2001. doi:10.1074/jbc.M106227200. PMID 11602582.
- ↑ "Crystallization of native striated-muscle actin". FEBS Letters 73 (1): 89–91. Jan 1977. doi:10.1016/0014-5793(77)80022-7. PMID 320040.
- ↑ "Multiple crystal structures of actin dimers and their implications for interactions in the actin filament". Acta Crystallographica Section D 64 (Pt 4): 454–465. Apr 2008. doi:10.1107/S0907444908003351. PMID 18391412.
- ↑ "Structural basis of actin filament capping at the barbed-end: a cryo-electron microscopy study". The EMBO Journal 25 (23): 5626–5633. Nov 2006. doi:10.1038/sj.emboj.7601395. PMID 17110933.
- ↑ 198.0 198.1 Lodish et al. 2016, pp. 791–792.
- ↑ 199.0 199.1 199.2 Lodish et al. 2016, pp. 792.
Works cited
- "Cell Organization and Movement I: Microfilaments". Molecular Cell Biology (Eighth ed.). New York: W.H. Freeman. 2016. ISBN 978-1-4641-8339-3.
- "Actin and Actin-Binding Proteins". Cold Spring Harb Perspect Biol 8 (8): a018226. August 2016. doi:10.1101/cshperspect.a018226. PMID 26988969.
External links
- Actin Staining Techniques (Live and Fixed Cell Staining)
- Eukaryotic Linear Motif resource motif class LIG_Actin_RPEL_3
- Eukaryotic Linear Motif resource motif class LIG_Actin_WH2_1
- Eukaryotic Linear Motif resource motif class LIG_Actin_WH2_2
- 3D macromolecular structures of actin filaments from the EM Data Bank(EMDB)
 KSF
KSF