Tyrosine aminotransferase
 From Wikipedia - Reading time: 9 min
From Wikipedia - Reading time: 9 min| Tyrosine transaminase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Human tyrosine aminotransferase (rainbow colored, N-terminus = blue, C-terminus = red) complexed with pyridoxal phosphate (space-filling model).[1] | |||||||||
| Identifiers | |||||||||
| EC no. | 2.6.1.5 | ||||||||
| CAS no. | 9014-55-5 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| TAT | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | TAT, Tat, tyrosine aminotransferase | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 613018 MGI: 98487 HomoloGene: 37293 GeneCards: TAT | ||||||||||||||||||||||||||||||||||||||||||||||||||
| EC number | 2.6.1.5 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Tyrosine aminotransferase (or tyrosine transaminase) is an enzyme present in the liver and catalyzes the conversion of tyrosine to 4-hydroxyphenylpyruvate.[6]

In humans, the tyrosine aminotransferase protein is encoded by the TAT gene.[7] A deficiency of the enzyme in humans can result in what is known as type II tyrosinemia, wherein there is an abundance of tyrosine as a result of tyrosine failing to undergo an aminotransferase reaction to form 4-hydroxyphenylpyruvate.[8]
Mechanism[edit]
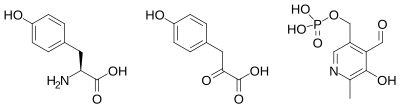
Structures of the three main molecules involved in chemical reaction catalyzed by the tyrosine aminotransferase enzyme are shown below: the amino acid tyrosine (left), the prosthetic group pyridoxal phosphate (right), and the resulting product 4-hydroxyphenylpyruvate (center).
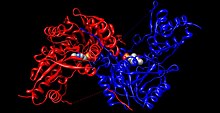
Each side of the dimer protein includes pyridoxal phosphate (PLP) bonded to the Lys280 residue of the tyrosine aminotransferase molecule. The amine group of tyrosine attacks the alpha carbon of the imine bonded to Lys280, forming a tetrahedral complex and then kicking off the LYS-ENZ. This process is known as transimination by the act of switching out the imine group bonded to PLP. The newly formed PLP-TYR molecule is then attacked by a base.
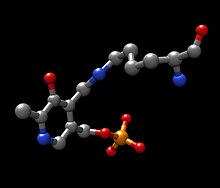
A possible candidate for the base in the mechanism could be Lys280 that was just pushed off of PLP, which sequesters the newly formed amino group of the PLP-TYR molecule. In a similar mechanism of aspartate transaminase, the lysine that forms the initial imine to PLP later acts as the base that attacks the tyrosine in transimination. The electrons left behind from the loss of the proton move down to form a new double bond to the imine, which in turn pushes the already double bonded electrons through PLP and end up as a lone pair on the positively charged nitrogen in the six-membered ring of the molecule. Water attacks the alpha carbon of the imine of PLP-TYR and through acyl substitution kicks off the nitrogen of PLP and forming pyridoxamine phosphate (PMP) and 4-hydroxyphenylpyruvate.

PMP is then regenerated into PLP by transferring its amine group to alpha-ketoglutarate, reforming its aldehyde functional group. This is followed by another substitution reaction with the Lys280 residue to reform its imine linkage to the enzyme, forming ENZ-PLP.
Active site[edit]

Tyrosine Aminotransferase as a dimer has two identical active site. Lys280 is attached to PLP, which is held in place via two nonpolar amino acid side chains; phenylalanine and isoleucine (see thumbnail on right). The PLP is also held in place by hydrogen bonding to surrounding molecules mainly by its phosphate group.
Shown below is one active site at three different magnifications:
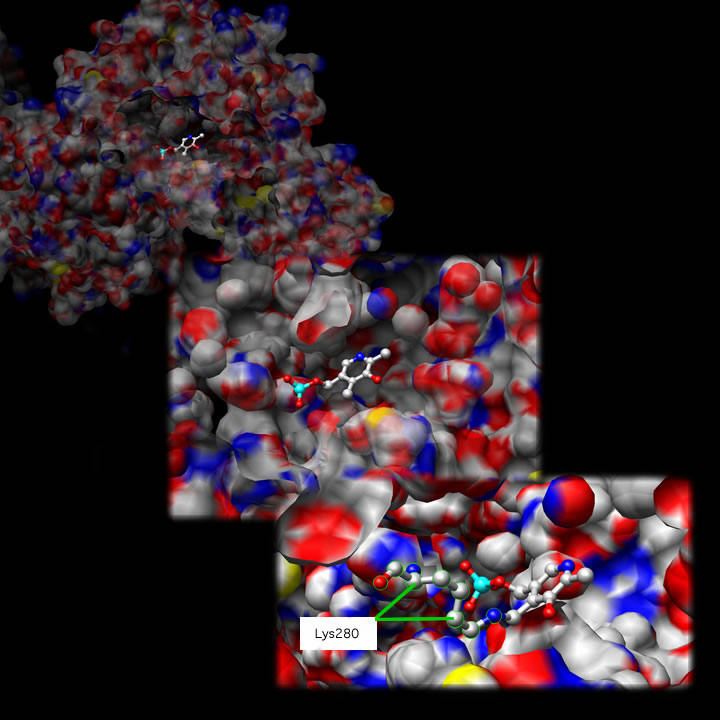

Pathology[edit]
Tyrosinemia is the most common metabolic disease associated with tyrosine aminotransferase. The disease results from a deficiency in hepatic tyrosine aminotransferase.[10] Tyrosinemia type II (Richner-Hanhart syndrome, RHS) is a disease of autosomal recessive inheritance characterized by keratitis, palmoplantar hyperkeratosis, mental retardation, and elevated blood tyrosine levels.[10] Keratitis in Tyrosinemia type II patients is caused by the deposition of tyrosine crystals in the cornea and results in corneal inflammation.[11] The TAT gene is located on human chromosome 16q22-24 and extends over 10.9 kilobases (kb) containing 12 exons, and its 3.0 kb mRNA codes for a 454-amino acid protein of 50.4 kDa.[12] Twelve different TAT gene mutations have been reported.[12]
References[edit]
- ^ a b PDB: 3DYD; Karlberg T, Moche M, Andersson J, et al. (2008). "Human tyrosine aminotransferase". Protein Data Bank. doi:10.2210/pdb3dyd/pdb.
- ^ a b c GRCh38: Ensembl release 89: ENSG00000198650 - Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000001670 - Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Dietrich JB (April 1992). "Tyrosine aminotransferase: a transaminase among others?". Cellular and Molecular Biology. 38 (2): 95–114. PMID 1349265.
- ^ Zea-Rey AV, Cruz-Camino H, Vazquez-Cantu DL, Gutiérrez-García VM, Santos-Guzmán J, Cantú-Reyna C (27 November 2017). "The Incidence of Transient Neonatal Tyrosinemia Within a Mexican Population". Journal of Inborn Errors of Metabolism and Screening. 5: 232640981774423. doi:10.1177/2326409817744230.
- ^ Rettenmeier R, Natt E, Zentgraf H, Scherer G (July 1990). "Isolation and characterization of the human tyrosine aminotransferase gene". Nucleic Acids Res. 18 (13): 3853–61. doi:10.1093/nar/18.13.3853. PMC 331086. PMID 1973834.
- ^ Pettersen, E.F., Goddard, T.D., Huang, C.C., Couch, G.S., Greenblatt, D.M., Meng, E.C., Ferrin, T.E. (2004). "UCSF Chimera - A Visualization System for Exploratory Research and Analysis". Journal of Computational Chemistry. 25 (13): 1605–1612. CiteSeerX 10.1.1.456.9442. doi:10.1002/jcc.20084. PMID 15264254. S2CID 8747218.
- ^ a b Natt E, Kida K, Odievre M, Di Rocco M, Scherer G (October 1992). "Point mutations in the tyrosine aminotransferase gene in tyrosinemia type II". Proc. Natl. Acad. Sci. U.S.A. 89 (19): 9297–301. Bibcode:1992PNAS...89.9297N. doi:10.1073/pnas.89.19.9297. PMC 50113. PMID 1357662.
- ^ al-Hemidan AI, al-Hazzaa SA (March 1995). "Richner-Hanhart syndrome (tyrosinemia type II). Case report and literature review". Ophthalmic Genet. 16 (1): 21–6. doi:10.3109/13816819509057850. PMID 7648039.
- ^ a b Minami-Hori M, Ishida-Yamamoto A, Katoh N, Takahashi H, Iizuka H (January 2006). "Richner-Hanhart syndrome: report of a case with a novel mutation of tyrosine aminotransferase". J. Dermatol. Sci. 41 (1): 82–4. doi:10.1016/j.jdermsci.2005.10.007. PMID 16318910.
Molecular graphics images were produced using the UCSF Chimera package from the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco (supported by NIH P41 RR-01081).
External links[edit]
- Tyrosine+aminotransferase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- PDBe-KB provides an overview of all the structure information available in the PDB for Human Tyrosine aminotransferase
Original source: https://en.wikipedia.org/wiki/Tyrosine aminotransferase
Status: article is cached
 KSF
KSF