Connexin
 From Wikipedia - Reading time: 14 min
From Wikipedia - Reading time: 14 min
| Connexin | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Connexin-26 dodecamer. A gap junction, composed of twelve identical connexin proteins, six in the membrane of each cell. Each of these six units is a single polypeptide which passes the membrane four times (referred to as four-pass transmembrane proteins). | |||||||||
| Identifiers | |||||||||
| Symbol | Connexin | ||||||||
| Pfam | PF00029 | ||||||||
| InterPro | IPR013092 | ||||||||
| PROSITE | PDOC00341 | ||||||||
| TCDB | 1.A.24 | ||||||||
| OPM superfamily | 194 | ||||||||
| OPM protein | 2zw3 | ||||||||
| |||||||||
Connexins (Cx) (TC# 1.A.24), or gap junction proteins, are structurally related transmembrane proteins that assemble to form vertebrate gap junctions. An entirely different family of proteins, the innexins, forms gap junctions in invertebrates.[1] Each gap junction is composed of two hemichannels, or connexons, which consist of homo- or heterohexameric arrays of connexins, and the connexon in one plasma membrane docks end-to-end with a connexon in the membrane of a closely opposed cell. The hemichannel is made of six connexin subunits, each of which consist of four transmembrane segments. Gap junctions are essential for many physiological processes, such as the coordinated depolarization of cardiac muscle, proper embryonic development, and the conducted response in microvasculature. Connexins also have non-channel dependant functions relating to cytoskeleton and cell migration.[2] For these reasons, mutations in connexin-encoding genes can lead to functional and developmental abnormalities.
Nomenclature
[edit]Connexins are commonly named according to their molecular weights, e.g. Cx26 is the connexin protein of 26 kDa. A competing nomenclature is the gap junction protein system, where connexins are sorted by their α (GJA) and β (GJB) forms, with additional connexins grouped into the C, D and E groupings, followed by an identifying number, e.g. GJA1 corresponds to Cx43. Following a vote at the Gap Junction Conference (2007) in Elsinore the community agreed to use the GJ nomenclature system for the genes that encode connexins, but wished to retain the connexin nomenclature for the encoded proteins using the weight of the human protein for the numbering of orthologous proteins.
Structure
[edit]
Connexins contain four highly ordered transmembrane segments (TMSs), primarily unstructured C and N cytoplasmic termini, a cytoplasmic loop (CL) and two extra-cellular loops, (EL-1) and (EL-2). Connexins are assembled in groups of six to form hemichannels, or connexons, and two hemichannels then combine to form a gap junction.
The crystal structure of the gap junction channel formed by human Cx26 (also known as GJB2) at 3.5 Å resolution is available.[3] The density map showed the two membrane-spanning hemichannels and the arrangement of the four TMSs of the six protomers forming each hemichannel. The hemichannels feature a positively charged cytoplasmic entrance, a funnel, a negatively charged transmembrane pathway, and an extracellular cavity. The pore is narrowed at the funnel, which is formed by the six amino-terminal helices lining the wall of the channel, which thus determines the molecular size restriction at the channel entrance.
The connexin gene family is diverse, with twenty-one identified members in the sequenced human genome, and twenty in the mouse (nineteen of which are orthologous pairs). They usually weigh between 25 and 60 kDa, and have an average length of 380 amino acids. The various connexins have been observed to combine into both homomeric and heteromeric gap junctions, each of which may exhibit different functional properties including pore conductance, size selectivity, charge selectivity, voltage gating, and chemical gating.[4]
Biosynthesis and internalization
[edit]A remarkable aspect of connexins is that they have a relatively short half life of only a few hours.[5] The result is the presence of a dynamic cycle by which connexins are synthesized and replaced. It has been suggested that this short life span allows for more finely regulated physiological processes to take place, such as in the myometrium.
From the nucleus to the membrane
[edit]As they are being translated by ribosomes, connexins are inserted into the membrane of the endoplasmic reticulum (ER).[6] It is in the ER that connexins are properly folded, yielding two extracellular loops, EL-1 and EL-2. It is also in the ER that the oligomerization of connexin molecules into hemichannels begins, a process which may continue in the UR-Golgi intermediate compartment as well.[5] The arrangements of these hemichannels can be homotypic, heterotypic, and combined heterotypic/heteromeric. After exiting the ER and passing through the ERGIC, the folded connexins will usually enter the cis-Golgi network.[7] However, some connexins, such as Cx26 may be transported independent of the Golgi.[8][9][10][11][12]
Gap junction assembly
[edit]After being inserted into the plasma membrane of the cell, the hemichannels freely diffuse within the lipid bilayer.[13] Through the aid of specific proteins, mainly cadherins, the hemichannels are able to dock with hemichannels of adjacent cells forming gap junctions.[14] Recent studies have shown the existence of communication between adherens junctions and gap junctions,[15] suggesting a higher level of coordination than previously thought.
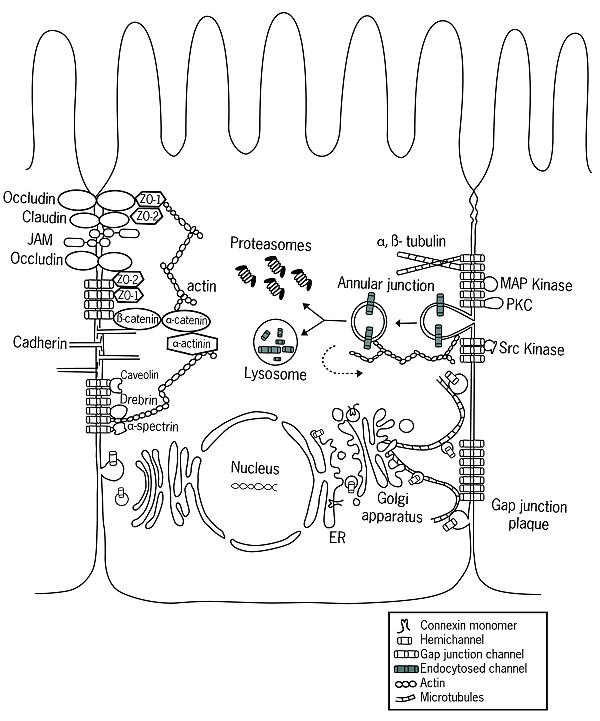
Function
[edit]Connexin gap junctions are found only in vertebrates, while a functionally analogous (but genetically unrelated) group of proteins, the innexins, are responsible for gap junctions in invertebrate species. Innexin orthologs have also been identified in Chordates, but they are no longer capable of forming gap junctions. Instead, the channels formed by these proteins (called pannexins) act as very large transmembrane pores that connect the intra- and extracellular compartments.
Within the CNS, gap junctions provide electrical coupling between progenitor cells, neurons, and glial cells. By using specific connexin knockout mice, studies revealed that cell coupling is essential for visual signaling. In the retina, ambient light levels influence cell coupling provided by gap junction channels, adapting the visual function for various lighting conditions. Cell coupling is governed by several mechanisms, including connexin expression.[17]
Decrock et al.. have discussed a multilevel platform via which connexins and pannexins can influence the following cellular functions within a tissue: (1) connexin gap junctional channels (GJCs) enable direct cell-cell communication of small molecules, (2) connexin hemichannels and pannexin channels can contribute to autocrine/paracrine signaling pathways, and (3) different structural domains of these proteins allow for channel-independent functions, such as cell-cell adhesion, interactions with the cytoskeleton, and the activation of intracellular signaling pathways.[18] Thus, connexins and pannexins have multifaceted contributions to brain development and specific processes in the neuro-glio-vascular unit, including synaptic transmission and plasticity, glial signaling, vasomotor control, cell movement, and blood-brain barrier integrity in the mature CNS.[18][2]
Substrate specificity
[edit]Different connexins may exhibit differing specificities for solutes. For example, adenosine passed about 12-fold better through channels formed by Cx32 while AMP and ADP passed about 8-fold better, and ATP greater than 300-fold better, through channels formed by Cx43. Thus, addition of phosphate to adenosine appears to shift its relative permeability from channels formed by Cx32 to channels formed by Cx43. This may have functional consequence because the energy status of a cell could be controlled via connexin expression and channel formation.[19]
Transport reaction
[edit]The transport reaction catalyzed by connexin gap junctions is:
- Small molecules (cell 1 cytoplasm) ⇌ small molecules (cell 2 cytoplasm)
Human connexins and clinical significance
[edit]| Connexin | Gene | Location and Function |
|---|---|---|
| Cx43 | GJA1 | Expressed at the surface of vasculature with atherosclerotic plaque, and up-regulated during atherosclerosis in mice. May have pathological effects. Also expressed between granulosa cells, which is required for proliferation. Normally expressed in astrocytes, also detected in most of the human astrocytomas and in the astroglial component of glioneuronal tumors.[20] It is also the main cardiac connexin, found mainly in ventricular myocardium.[21] Associated with oculodentodigital dysplasia. |
| Cx46 | GJA3 | |
| Cx37 | GJA4 | Induced in vascular smooth muscle during coronary arteriogenesis. Cx37 mutations are not lethal. Forms gap junctions between oocytes and granulosa cells, and are required for oocyte survival. |
| Cx40 | GJA5 | Expressed selectively in atrial myocytes. Responsible for mediating the coordinated electrical activation of atria.[22] |
| Cx33 | GJA6 (GJA6P) |
Pseudogene in humans |
| Cx50 | GJA8 | Gap junctions between A-typ horizontal cells in mouse and rabbit retina[23] |
| Cx59 | GJA10 | |
| Cx62 | GJA10 | Human Cx62 complies Cx57 (mouse). Location in axon-bearing B-typ horizontal cell in rabbit retina[24] |
| Cx32 | GJB1 | Major component of the peripheral myelin. Mutations in the human gene cause X-linked Charcot-Marie-Tooth disease, a hereditary neuropathy. In human normal brain CX32 expressed in neurons and oligodendrocytes.[20] |
| Cx26 | GJB2 | Mutated in Vohwinkel syndrome[25] as well as Keratitis-Icthyosis-Deafness (KID) Syndrome.[25] |
| Cx31 | GJB3 | Can be associated with Erythrokeratodermia variabilis. |
| Cx30.3 | GJB4 | Fonseca et al. confirmed Cx30.3 expression in thymocytes.[26] Can be associated with Erythrokeratodermia variabilis. |
| Cx31.1 | GJB5 | |
| Cx30 | GJB6 | Mutated in Clouston syndrome (hidrotic ectodermal dysplasia)[25] |
| Cx25 | GJB7 | |
| Cx45 | GJC1/GJA7 | Human pancreatic ductal epithelial cells.[27] Atrio-ventricular node. |
| Cx47 | GJC2/GJA12 | Expressed in oligodendrocyte gap junctions[28] |
| Cx31.3 | GJC3 | Human ortholog of murine Cx29. Not known to form gap junctions.[29] |
| Cx36 | GJD2/GJA9 | Pancreatic beta cell function, mediating the release of insulin. Neurons throughout the central nervous system where they synchronize neural activity.[30] |
| Cx31.9 | GJD3/GJC1 | |
| Cx39 | GJD4 | |
| Cx40.1 | GJD4 | |
| Cx23 | GJE1 |
Gap junctions are essential for many physiological processes, such as the coordinated depolarization of cardiac muscle, proper embryonic development, and the conducted response in microvasculature. For this reason, deletion or mutation of the various connexin isoforms produces distinctive phenotypes and pathologies.[31] While mutations in Cx43 are mostly linked to oculodentodigital dysplasia, Cx47 mutations are associated with Pelizaeus-Merzbacher-like disease and lymphedema. Cx40 mutations are principally linked to atrial fibrillation. Mutations in Cx37 have not yet been described, but polymorphisms in the Cx37 gene have been implicated in the development of arterial disease.[32][33]
References
[edit]- ^ Lodish HF, Berk A, Matsudaira P, Kaiser CA, Krieger M, Scott MP, Zipursky SL, Darnell J (2004). Molecular Cell Biology (5th ed.). New York: W.H. Freeman and Company. pp. 230–31. ISBN 0-7167-4366-3.
- ^ a b Matsuuchi L, Naus CC (January 2013). "Gap junction proteins on the move: connexins, the cytoskeleton and migration". Biochim Biophys Acta. 1828 (1): 94–108. doi:10.1016/j.bbamem.2012.05.014. PMID 22613178.
- ^ Maeda S, Nakagawa S, Suga M, Yamashita E, Oshima A, Fujiyoshi Y, Tsukihara T (April 2009). "Structure of the connexin 26 gap junction channel at 3.5 A resolution". Nature. 458 (7238): 597–602. Bibcode:2009Natur.458..597M. doi:10.1038/nature07869. ISSN 1476-4687. PMID 19340074. S2CID 4431769.
- ^ Ayad WA, Locke D, Koreen IV, Harris AL (June 2006). "Heteromeric, but not homomeric, connexin channels are selectively permeable to inositol phosphates". J. Biol. Chem. 281 (24): 16727–39. doi:10.1074/jbc.M600136200. ISSN 0021-9258. PMID 16601118.
- ^ a b Laird DW (March 2006). "Life cycle of connexins in health and disease". Biochem. J. 394 (Pt 3): 527–43. doi:10.1042/BJ20051922. PMC 1383703. PMID 16492141.
- ^ Bennett MV, Zukin RS (February 2004). "Electrical coupling and neuronal synchronization in the Mammalian brain". Neuron. 41 (4): 495–511. doi:10.1016/s0896-6273(04)00043-1. PMID 14980200. S2CID 18566176.
- ^ Musil LS, Goodenough DA (September 1993). "Multisubunit assembly of an integral plasma membrane channel protein, gap junction connexin43, occurs after exit from the ER". Cell. 74 (6): 1065–77. doi:10.1016/0092-8674(93)90728-9. PMID 7691412. S2CID 12169415.
- ^ Evans WH, Ahmad S, Diez J, George CH, Kendall JM, Martin PE (1999). "Trafficking pathways leading to the formation of gap junctions". Novartis Foundation Symposium 219 - Gap Junction-Mediated Intercellular Signalling in Health and Disease. Novartis Foundation Symposia. Vol. 219. pp. 44–54, discussion 54–9. doi:10.1002/9780470515587.ch4. ISBN 9780470515587. PMID 10207897.
- ^ George CH, Kendall JM, Evans WH (March 1999). "Intracellular trafficking pathways in the assembly of connexins into gap junctions". J. Biol. Chem. 274 (13): 8678–85. doi:10.1074/jbc.274.13.8678. PMID 10085106.
- ^ George CH, Kendall JM, Campbell AK, Evans WH (November 1998). "Connexin-aequorin chimerae report cytoplasmic calcium environments along trafficking pathways leading to gap junction biogenesis in living COS-7 cells". J. Biol. Chem. 273 (45): 29822–9. doi:10.1074/jbc.273.45.29822. PMID 9792698.
- ^ Martin PE, George CH, Castro C, Kendall JM, Capel J, Campbell AK, Revilla A, Barrio LC, Evans WH (January 1998). "Assembly of chimeric connexin-aequorin proteins into functional gap junction channels. Reporting intracellular and plasma membrane calcium environments". J. Biol. Chem. 273 (3): 1719–26. doi:10.1074/jbc.273.3.1719. PMID 9430718.
- ^ Martin PE, Errington RJ, Evans WH (2001). "Gap junction assembly: multiple connexin fluorophores identify complex trafficking pathways". Cell Commun. Adhes. 8 (4–6): 243–8. doi:10.3109/15419060109080731. PMID 12064596. S2CID 3029281.
- ^ Thomas T, Jordan K, Simek J, Shao Q, Jedeszko C, Walton P, Laird DW (October 2005). "Mechanisms of Cx43 and Cx26 transport to the plasma membrane and gap junction regeneration". J. Cell Sci. 118 (Pt 19): 4451–62. doi:10.1242/jcs.02569. PMID 16159960. S2CID 13486416.
- ^ Jongen WM, Fitzgerald DJ, Asamoto M, Piccoli C, Slaga TJ, Gros D, Takeichi M, Yamasaki H (August 1991). "Regulation of connexin 43-mediated gap junctional intercellular communication by Ca2+ in mouse epidermal cells is controlled by E-cadherin". J. Cell Biol. 114 (3): 545–55. doi:10.1083/jcb.114.3.545. PMC 2289094. PMID 1650371.
- ^ Wei CJ, Francis R, Xu X, Lo CW (May 2005). "Connexin43 associated with an N-cadherin-containing multiprotein complex is required for gap junction formation in NIH3T3 cells" (PDF). J. Biol. Chem. 280 (20): 19925–36. doi:10.1074/jbc.M412921200. PMID 15741167. S2CID 770387.
- ^ Dbouk HA, Mroue RM, El-Sabban ME, Talhouk RS (March 2009). "Connexins: a myriad of functions extending beyond assembly of gap junction channels". Cell Commun Signal. 7: 4. doi:10.1186/1478-811X-7-4. PMC 2660342. PMID 19284610.
- ^ Kihara AH, de Castro LM, Moriscot AS, Hamassaki DE (May 2006). "Prolonged dark adaptation changes connexin expression in the mouse retina". J Neurosci Res. 83 (7): 1331–41. doi:10.1002/jnr.20815. PMID 16496335. S2CID 2919282.
- ^ a b Decrock E, De Bock M, Wang N, Bultynck G, Giaume C, Naus CC, Green CR, Leybaert L (August 2015). "Connexin and pannexin signaling pathways, an architectural blueprint for CNS physiology and pathology?". Cell. Mol. Life Sci. 72 (15): 2823–51. doi:10.1007/s00018-015-1962-7. ISSN 1420-9071. PMC 11113968. PMID 26118660. S2CID 17170098.
- ^ Goldberg GS, Moreno AP, Lampe PD (September 2002). "Gap junctions between cells expressing connexin 43 or 32 show inverse permselectivity to adenosine and ATP". J. Biol. Chem. 277 (39): 36725–30. doi:10.1074/jbc.M109797200. ISSN 0021-9258. PMID 12119284.
- ^ a b Aronica E, Gorter JA, Jansen GH, Leenstra S, Yankaya B, Troost D (May 2001). "Expression of connexin 43 and connexin 32 gap-junction proteins in epilepsy-associated brain tumors and in the perilesional epileptic cortex". Acta Neuropathol. 101 (5): 449–59. doi:10.1007/s004010000305. PMID 11484816. S2CID 6738913.
- ^ Verheule S, van Kempen MJ, te Welscher PH, Kwak BR, Jongsma HJ (May 1997). "Characterization of gap junction channels in adult rabbit atrial and ventricular myocardium". Circ. Res. 80 (5): 673–81. doi:10.1161/01.res.80.5.673. PMID 9130448.
- ^ Gollob MH, Jones DL, Krahn AD, Danis L, Gong XQ, Shao Q, et al. (June 2006). "Somatic mutations in the connexin 40 gene (GJA5) in atrial fibrillation". N. Engl. J. Med. 354 (25): 2677–88. doi:10.1056/NEJMoa052800. PMID 16790700.
- ^ Massey, Stephen (16 January 2009). Connexins: A Guide (1st ed.). Springer-Verlag Gmbh. pp. 3–?. ISBN 978-1-934115-46-6.
- ^ Beyer, Eric C.; Berthound, Viviana M. (16 January 2009). Connexins: A Guide (1st ed.). Springer-Verlag Gmbh. pp. 387–417. ISBN 978-1-934115-46-6.
- ^ a b c Avshalumova L, Fabrikant J, Koriakos A (February 2014). "Overview of skin diseases linked to connexin gene mutations". Int J Dermatol. 53 (2): 192–205. doi:10.1111/ijd.12062. PMID 23675785. S2CID 205187359.
- ^ Fonseca PC, Nihei OK, Urban-Maldonado M, Abreu S, de Carvalho AC, Spray DC, Savino W, Alves LA (June 2004). "Characterization of connexin 30.3 and 43 in thymocytes". Immunol. Lett. 94 (1–2): 65–75. doi:10.1016/j.imlet.2004.03.019. PMID 15234537.
- ^ Tai MH, Olson LK, Madhukar BV, Linning KD, Van Camp L, Tsao MS, Trosko JE (January 2003). "Characterization of gap junctional intercellular communication in immortalized human pancreatic ductal epithelial cells with stem cell characteristics". Pancreas. 26 (1): e18–26. doi:10.1097/00006676-200301000-00025. PMID 12499933. S2CID 34571252.
- ^ Kamasawa N, Sik A, Morita M, Yasumura T, Davidson KG, Nagy JI, Rash JE (2005). "Connexin-47 and connexin-32 in gap junctions of oligodendrocyte somata, myelin sheaths, paranodal loops and Schmidt-Lanterman incisures: implications for ionic homeostasis and potassium siphoning". Neuroscience. 136 (1): 65–86. doi:10.1016/j.neuroscience.2005.08.027. PMC 1550704. PMID 16203097.
- ^ Sargiannidou I, Ahn M, Enriquez AD, Peinado A, Reynolds R, Abrams C, Scherer SS, Kleopa KA (May 2008). "Human oligodendrocytes express Cx31.3: function and interactions with Cx32 mutants". Neurobiol. Dis. 30 (2): 221–33. doi:10.1016/j.nbd.2008.01.009. PMC 2704064. PMID 18353664.
- ^ Connors BW, Long MA (2004). "Electrical synapses in the mammalian brain". Annu. Rev. Neurosci. 27: 393–418. doi:10.1146/annurev.neuro.26.041002.131128. PMID 15217338.
- ^ Pfenniger A, Wohlwend A, Kwak BR (January 2011). "Mutations in connexin genes and disease". Eur. J. Clin. Invest. 41 (1): 103–16. doi:10.1111/j.1365-2362.2010.02378.x. ISSN 1365-2362. PMID 20840374. S2CID 24404442.
- ^ Fang JS, Burt JM (September 2022). "Connexin37 Regulates Cell Cycle in the Vasculature". J Vasc Res. 60 (2): 73–86. doi:10.1159/000525619. PMID 36067749.
- ^ Molica F, Meens MJ, Morel S, Kwak BR (September 2014). "Mutations in cardiovascular connexin genes". Biology of the Cell. 106 (9): 269–93. doi:10.1111/boc.201400038. PMID 24966059. S2CID 10070999.
Sources
[edit]- As of this edit, this article uses content from "1.A.24 The Gap Junction-forming Connexin (Connexin) Family", which is licensed in a way that permits reuse under the Creative Commons Attribution-ShareAlike 3.0 Unported License, but not under the GFDL. All relevant terms must be followed.
External links
[edit] Media related to connexins at Wikimedia Commons
Media related to connexins at Wikimedia Commons- Connexins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
 KSF
KSF