Genetic history of East Asians
 From Wikipedia - Reading time: 58 min
From Wikipedia - Reading time: 58 min
This article summarizes the genetic makeup and population history of East Asian peoples and their connection to genetically related populations such as Southeast Asians and North Asians, as well as Oceanians, and partly, Central Asians, South Asians, and Native Americans. They are collectively referred to as "East Eurasians" in population genomics.[1]
Overview
[edit]Origins
[edit]-
Phylogenetic position of East Asian lineages among other Eastern Eurasians
-
Schematic of Populations in Eurasia from 45 to 10 kaBP
-
Highlighted regions show where ancient individuals associated with the labeled ancestry have been sampled

Population genomic research has studied the origin and formation of modern East Asians. The ancestors of East Asians (Ancient East Eurasians) split from other human populations (Ancient West Eurasians) possibly as early as 80,000 to 50,000 years ago. Possible routes into East Asia include a northern route from Central Asia, beginning north of the Himalayas, and a southern route, beginning south of the Himalayas and moving through South and Southeast Asia.[3][4][1][5] Vallini et al. 2024 noted that the divergence between Ancient East Eurasians and West Eurasians most likely occurred on the Persian Plateau between 57–46kya years ago.[6]
Phylogenetic data suggests that an early Initial Upper Paleolithic wave (earlier than 45kya) "ascribed to a population movement with uniform genetic features and material culture" (Ancient East Eurasians) dispersed throughout Eurasia, with one of its main branches (dubbed as East Eurasian Core; EEC) using a Southern dispersal route through Southern Asia, where they subsequently diverged rapidly into the ancestors of Australasians (Oceanians), the Ancient Ancestral South Indians (AASI), as well as Andamanese and East/Southeast Asians (ESEA),[7][4][1][8] although Papuans may have also received some geneflow from an earlier group (xOoA),[9] around 2%,[10] next to additional archaic admixture in the Sahul region.

Deeply diverged Ancient East Eurasian lineages associated with the spread of IUP-affilated material culture along an early 'northern route' into eastern Europe (Bacho Kiro Cave and Peștera cu Oase), Central Asia (Ust'-Ishim man), and Siberia (Kara-Bom etc.), as well as Northwest China, did contribute only little or no ancestry to modern East Asians, which instead were found to descend primarily from the southern route wave.[12][13][14][15][8][16][1][4][6]
The single southern route dispersal into the South-Southeast Asia region gave rise to the AASI, Andamanese, Eastern Asian and Australasian populations. The southern route origin is strongly supported by archaeogenetic data and genetic affinities between these "East Eurasian Core" (EEC) populations. Although the morphological more distinct traits of modern Northeast Asian and Siberian populations raised the question of a possible distinct northern origin, those traits have subsequently been associated with adaptions to the extremely cold climate during the Last Glacial Maximum (LGM) of 24– 16 kya by an ancient people with initially southern affinities, which moved into Northeast Asia and Siberia from Southeast Asia. This is further supported by higher morphological affinities of ancient East Asian specimens, such as the Tianyuan man, the Zhoukoudian Upper Cave remains, the Liujiang man, the Red Deer Cave people, the Jōmon people as well as the Liangdao and Qihe Cave remains to "southern" populations and ancient remains, such as the Niah cave and Wajak remains or Hoabinhians, as well as modern Andamanese, Vedda, and Aboriginal Australians, but being genetically closer or basal to the derived Northeast Asian and Siberian groups.[8][1][17][18][19][20][21]
Additionally, genetic diversity within present-day Asian populations, follows a strong correlation with latitude: genetic diversity is decreasing from south to north. The correlation continues to hold true when only mainland Southeast Asian and East Asian populations are considered. This may be attributable to a serial founder effect, and is consistent with a single eastward migration of modern humans along a southern route into South and Southeast Asia. Subsequently, ancestral East Asians diversified in southern East Asia and subsequently dispersed northward across the continent.[8][1][17][18][19][20][21] The dominant paternal haplogroups of East Asians associated with the "southern route" belong to subclades of C-M217, D-M174, O-M175, and N-M231, among some others.[22]
The exact patterns of the northwards dispersal of ancestral East Asians from Southeast Asia remains unclear. There may have been both an "interior route" and a "coastal route", correlating in part with the distribution and frequency of paternal haplogroup subclades.[23][1][24] Accordingly, ancient and modern East Asians can be modeled as admixture between both a deeply branching interior lineage (represented by the Tianyuan specimen) and an equally deep branching coastal lineage (represented by present-day Önge peoples). The amounts of interior and coastal contributions vary depending on each model; while one study estimated 90% interior + 10% coastal contributions,[23] others estimated between 21–26% interior + 74–79% coastal contributions,[24] or nearly equal amounts of interior and coastal contributions (51–56% and 44–49% respectively).[25][26] There are also alternative scenarios for the divergence patterns of ancient East Asians without a distinct interior/coastal dispersal.[1][17]

There is also genetic evidence for migrations to Northern Asia and Siberia from both a deeply diverged East Eurasian Initial Upper Paleolithic group (related to the Ust'-Ishim man and or the Bacho Kiro cave remains) via Central Asia, which may have contributed ancestry (up to 39%[27]) to the Tianyuan lineage, and from a later Upper Paleolithic West Eurasian-affilated source, which contributed to the formation of Ancient North Eurasians (ANE/ANS). While the southern migration wave likely diversified after settling within East Asia, the wave associated with Upper Paleolithic Europeans mixed with the southern wave somewhere in Siberia. The ANE derive around 2/3 (50–71%) of their ancestry from a West Eurasian-like source best represented by Upper Paleolithic Europeans, and around 1/3 (29–50%) from an East Eurasian source best represented by the Tianyuan man or Upper Paleolithic East/Southeast Asians.[4][28][29][30] The legacy of this Paleolithic admixture event is also evident by the later dispersal of haplogroups Q and R, as well as ANE-like ancestry throughout Northern Eurasia, but which had only limited influence on modern East Asian groups.[22][31][17][32][4]
Although Huang et al. (2021) found evidence for light skin being selected among the ancestral populations of West Eurasians and East Eurasians, prior to their divergence,[33] the main period for the selection of light skin alleles (such as rs1800414-G) started around 25,000 to 30,000 years ago in East Asia, after the northwards expansion from South Asia. The affiliated alleles are distinct from those observed among European/Middle Eastern populations.[34]
Diversification and substructure
[edit]After the peopling of the South and Southeast Asia region by the East Eurasian Core (EEC), this ancestral population diverged rapidly into at least three "deeply branching East Asian lineages, namely the "Ancient Ancestral South Indians" (AASI) staying in Southern Asia, Australasians (AA) heading into Oceania, and the East- and Southeast Asian (ESEA) branch in Southeast Asia and subsequently expanding northwards. The ESEA branch would become broadly ancestral to modern East Asians, but also Southeast Asians, Polynesians, and Siberians, as well as Native Americans. The ESEA source population is estimated to have expanded outgoing from Mainland Southeast Asia at c. 40,000 BCE. The deepest split among the ESEA lineage gave rise to the "basal Asian" Hoabinhian hunter-gatherers of Southeast Asia and the c. 39,000-year-old Tianyuan lineage in Northern China and the Amur Region, followed by the divergence of the Longlin lineage in Guangxi and the Jōmon lineages on the Japanese archipelago, and finally the split between Ancient Southern East Asians (ASEA) and Ancient Northern East Asians (ANEA) somewhere in East Asia. The divergence between ASEA and ANEA happened sometimes between 26,000 and 20,000 years ago.[7] ANEA ancestry replaced the earlier Tianyuan-like ancestry in Northern China, the Amur region and parts of Siberia since at least 19,000 years ago, while Longlin/Guangxi ancestry was largely replaced by ASEA ancestry since at least 9,000 years ago in Southern China.[35][1]
There are currently eight identified closely related sub-ancestries on the ESEA branch:[17][1]
- Hoabinhian ancestry – ancestry on the ESEA lineage associated with 8,000–4,000-year-old hunter-gatherers in Laos and Malaysia.
- Tianyuan ancestry – ancestry on the ESEA lineage associated with an Upper Paleolithic individual dating to c. 39,000 years ago in Northern China.
- Guangxi ancestry – associated with a 10,500-year-old individual from Longlin, Guangxi. This ancestry was not observed in either historical samples from Guangxi or contemporary East and Southeast Asians, suggesting that the lineage went largely extinct.
- Jōmon ancestry – ancestry associated with 8,000–3,000-year-old individuals in the Japanese archipelago.
- Ancient Northern East Asian (ANEA) – associated with populations in the Amur River region, Mongolia, Siberia, as well as in the Yellow River region to central China; ancestral to the derived Ancient Northeast Asian (ANA/Amur) ancestry. Sister lineage to ASEA.
- Ancient Southern East Asian (ASEA) – Associated with ancient samples in the Fujian and Guangxi region of Southern China. Sister lineage to ANEA.
- Yellow River ancestry – a sub-group of Ancient Northern East Asian, associated with a sample of a 9,500-year-old individual from the lower reaches of the Yellow River in Shandong, i.e. Bianbian. Formed either as admixture between primarily ANEA and minor ASEA, or as earlier admixture between a deep interior group (90%) and a deep coastal group (10%). Distinct from ANA, but branching deeply/basal within ANEA.
- Early Ancient Tibetan ancestry (EAT) – associated with the c. 6,000-year-old Zongri5.1K individual in the Himalayan region of the Tibetan Plateau. This ancestry is less present among modern populations, however it is still highest among the Qiang, Tibetan, and Sherpa people.

The genetic makeup of contemporary East Asians, such as the Han Chinese, is primarily characterized by the presence of "Yellow River" ancestry which formed from a major Ancient Northern East Asian (ANEA) component and a minor Ancient Southern East Asian (ASEA) one.[37][7]
Northeast Asians such as Tungusic, Mongolic, and Turkic peoples derive most of their ancestry from the "Amur" (Ancient Northeast Asian) subgroup of the Ancient Northern East Asians, which expanded massively with millet cultivation and pastoralism. Tungusic peoples display the highest genetic affinity to Ancient Northeast Asians, represented by c. 7,000 and 13,000 year old specimens, whereas Turkic-speaking peoples have variable but significant amounts of West Eurasian admixture.[38][35]
An early branch of Ancient Northern East Asians absorbed an Ancient North Eurasian population to their North, giving rise to the Ancient Paleo-Siberians, who in turn became ancestral to both "modern Paleo-Siberians" (such as Chukotko-Kamchatkan, Yeniseian, and Nivkh speakers) and contemporary Native Americans. Paleo-Siberian (APS) ancestry was once widespread across North Asia, but largely replaced by later waves of Neo-Siberian ancestry due to a major population turnover from the south, possibly involving Uralic and Yukaghir speakers. This was later followed by another expansion from the south in relatively recent times, associated with Amur ancestry (ANA) involving Tungusic, Mongolic, and Turkic speakers.[39][40]
Austronesians and Kra-Dai speakers in Southeast Asia mainly carry "Fujian Neolithic" ancestry, a subgroup of Ancient Southern East Asians, which is associated with the spread of rice cultivation. Isolated hunter-gatherers in Southeast Asia, specifically in Malaysia and Thailand, such as the Semang, derive most of their ancestry from the Hoabinhian lineage.[41][42][43] The emergence of the Neolithic in Southeast Asia went along with a population shift caused by migrations from Southern China. Neolithic Mainland Southeast Asian samples mainly have Ancient Southern East Asian ancestry (a sister lineage of "Fujian Neolithic" ancestry) with variable amounts of Hoabinhian-related admixture. In modern populations, this admixture of Ancient Southern East Asian and Hoabinhian ancestry is most strongly associated with Austroasiatic speakers.[44]
Studies on ancient and historical populations
[edit]Xiongnu people
[edit]The Xiongnu, possibly a Turkic, Mongolic, Yeniseian or multi-ethnic group, was a confederation[45] of nomadic peoples who, according to ancient Chinese sources, inhabited the eastern Eurasian Steppe from the 3rd century BC to the late 1st century AD. Chinese sources report that Modu Chanyu, the supreme leader after 209 BC, founded the Xiongnu Empire.[46]
Autosomal DNA
[edit]It was found that the "predominant part of the Xiongnu population is likely to have spoken Turkic". However, important cultural, technological and political elements may have been transmitted by Eastern Iranian-speaking Steppe nomads: "Arguably, these Iranian-speaking groups were assimilated over time by the predominant Turkic-speaking part of the Xiongnu population".[47] This is reflected by the average genetic makeup of Xiongnu samples, having approximately 58% East Eurasian ancestry, represented by a Bronze Age population from Khövsgöl, Mongolia, which may be associated with the Turkic linguistic heritage. The rest of the Xiongnu's ancestry (~40%) was related to West Eurasians, represented by the Gonur Depe BMAC population of Central Asia, and the Sintashta culture of the Western steppe.[47][48] The Xiongnu displayed striking heterogeneity and could be differentiated into two subgroups, "Western Xiongnu" and "Eastern Xiongnu", with the former being of "hybrid" origins displaying affinity to previous Saka tribes, such as represented by the Chandman culture, while the later was of primarily Ancient Northeast Asian (Ulaanzuukh-Slab Grave) origin.[49][47] High status Xiongnu individuals tended to have less genetic diversity, and their ancestry was essentially derived from the Eastern Eurasian Ulaanzuukh/Slab Grave culture.[50]
Paternal lineages
[edit]A review of the available research has shown that, as a whole, 53% of Xiongnu paternal haplogroups were East Eurasian, while 47% were West Eurasian.[51] In 2012, Chinese researchers published an analysis of the paternal haplogroups of 12 elite Xiongnu male specimens from Heigouliang in Xinjiang, China. Six of the specimens belonged to Q1a, while four belonged to Q1b-M378. 2 belonged to unidentified clades of Q*.[52] In another study, a probable Chanyu of the Xiongnu empire was assigned to haplogroup R1.[53][54]
Maternal lineages
[edit]The bulk of the genetics research indicates that, as a whole, 73% of Xiongnu maternal haplogroups were East Eurasian, while 27% were West Eurasian.[55] A 2003 study found that 89% of Xiongnu maternal lineages from the Egiin Gol valley were of East Asian origin, while 11% were of West Eurasian origin.[56] A 2016 study of Xiongnu from central Mongolia found a considerably higher frequency of West Eurasian maternal lineages, at 37.5%.[57]
Xianbei people
[edit]Autosomal DNA
[edit]A full genomic analysis performed on multiple Xianbei remains found the population to have derived primarily from the Ancient Northeast Asian gene pool.[58]
Paternal lineages
[edit]A genetic study published in the American Journal of Physical Anthropology in August 2018 noted that the paternal haplogroup C2b1a1b has been detected among the Xianbei and the Rouran, and was probably an important lineage among the Donghu people.[59]
Maternal lineages
[edit]Genetic studies published in 2006 and 2015 revealed that the mitochondrial haplogroups of Xianbei remains were of East Asian origin. According to Zhou (2006) the maternal haplogroup frequencies of the Tuoba Xianbei were 43.75% haplogroup D, 31.25% haplogroup C, 12.5% haplogroup B, 6.25% haplogroup A and 6.25% "other".[60] Zhou (2014) obtained mitochondrial DNA analysis from 17 Tuoba Xianbei, which indicated that these specimens were, similarly, completely East Asian in their maternal origins, belonging to haplogroups D, C, B, A, O and haplogroup G.[61][62]
Jōmon people
[edit]The Jōmon people represent the indigenous population of the Japanese archipelago during the Jōmon period. They are inferred to descend from the Paleolithic inhabitants of Japan. Genetic analyses on Jōmon remains found them to represent a deeply diverged East Asian lineage. The Jōmon lineage is inferred to have diverged from Ancient East Asians before the divergence between Ancient Northern East Asians and Ancient Southern East Asians, but after the divergence of the basal Tianyuan man and or Hoabinhians. Beyond their broad affinity with Eastern Asian lineages, the Jōmon also display a weak affinity for Ancient North Eurasians (ANE), which may be associated with the introduction of microblade technology to Northeast Asia and Northern East Asia during the Last Glacial Maximum via the ANE or Ancient Paleo-Siberians.[63][64]
Hoabinhians
[edit]The Hoabinhians represent a technologically advanced society of hunter-gatherers, primarily living in Mainland Southeast Asia, but also adjacent regions of Southern China. While the Upper Paleolithic origins of this 'Hoabinhian ancestry' are unknown, Hoabinhian ancestry has been found to be related to the main 'East Asian' ancestry component found in most modern East and Southeast Asians, although deeply diverged from it.[24][65] Together with the Paleolithic Tianyuan man, they form early branches of East Asian genetic diversity, and are described as "Basal Asian" (BA) or "Basal East Asian" (BEA).[66]
Studies on modern populations
[edit]
Manchu and Daur peoples
[edit]Autosomal DNA
[edit]A study on the Manchu population of Liaoning reported that they have a close genetic relationship and significant admixture signals from Northern Han Chinese. The Liaoning Manchu were formed from a major ancestral component related to Yellow River farmers and a minor ancestral component linked to ancient populations from the Amur River Basin, or others. The Manchu were therefore an exception to the coherent genetic structure of Tungusic-speaking populations, likely due to the large-scale population migrations and genetic admixtures in the past centuries.[68]
Paternal lineages
[edit]A plurality of Daur males belong to Haplogroup C-M217 (12/39 = 30.8% according to Xue Yali et al. 2006,[69] 88/207 = 42.5% according to Wang Chi-zao et al. 2018[70]), with Haplogroup O-M122 being the second most common haplogroup among present-day Daurs (10/39 = 25.6%,[69] 52/207 = 25.1%[70]). There are also tribes (hala; cf. Kazakh tribes) among the Daurs that belong predominantly to other Y-DNA haplogroups, such as Haplogroup N-M46/M178 (Merden hala) and Haplogroup O1b1a1a-M95 (Gobulo hala).[70] Haplogroup C3b2b1*-M401(xF5483)[71][72][73] has been identified as a possible marker of the Aisin Gioro and is found in ten different ethnic minorities in Northern China, but is less prevalent from Han Chinese.[74][75][73] The Manchu people also display a significant amount of haplogroup C-M217, but the most often observed Y-DNA haplogroup among present-day Manchus is Haplogroup O-M122, which they share in common with the general population of China.[76][69][77]
Ainu people
[edit]The exact origins of the early Ainu remains unclear, but it is generally agreed to be linked to the Satsumon culture of the Epi-Jōmon period, with later influences from the nearby Okhotsk culture.[78] The Ainu appear genetically most closely related to the Jōmon period peoples of Japan. The genetic makeup of the Ainu represents a "deep branch of East Asian diversity". Compared to contemporary East Asian populations, the Ainu share "a closer genetic relationship with northeast Siberians".[12][64]
Japanese people
[edit]
Japanese populations in modern Japan can be traced to three separate, but related demographics: the Ainu, Ryukyuan and Mainland Japanese (Yamato). The populations are closely related to clusters found in Northeastern Asia[81][82][83] with the Ainu group being most similar to the Ryukyuan group,[82][84] the Ryukyuan group being most similar to the Yamato group,[85] and the Yamato group being most similar to Koreans[86][87][88][89] among other East Asian people.
Autosomal DNA
[edit]The majority of Japanese genetic ancestry is derived from sources related to other mainland Asian groups, mostly Koreans, while the other amount is derived from the local Jōmon hunter-gatherers (9% ±3%).[90][91][89] Evidence for both Northern and Southern mtDNA and Y-DNA haplogroups has been observed in the Japanese, with the North-Eastern DNA taking up majority of the genetic makeup,[81] especially among the Mainland group.[87] In addition to the Northeastern ancestry, the Japanese demographics (alongside the Koreans), are the only ethnicities to have restricted presence of the Jōmon-like M7a DNA [ja] in East Asia.[92][93][67]
Paternal lineages
[edit]A comprehensive study of worldwide Y-DNA diversity (Underhill et al. 2000) included a sample of 23 males from Japan, of whom 35% belonged to haplogroup D-M174, 26% belonged to O-M175, 22% belonged to O-M122, 13% belonged to C-M8 and C-M130, and 4.3% belonged to N-M128.[94] Poznik et al. (2016) reported the haplogroups of a sample of Japanese men from Tokyo:[95] 36% belonged to D2-M179, 32% had O2b-M176, 18% carried O3-M122, 7.1% carried C1a1-M8, 3.6% belonged to O2a-K18, and 3.6% carried C2-M217.[96]
Maternal lineages
[edit]According to an analysis of the 1000 Genomes Project's sample of Japanese collected in the Tokyo metropolitan area, the mtDNA haplogroups found among modern Japanese include D (35.6%), B (13.6%), M7 (10.2%), G (10.2%), N9 (8.5%), F (7.6%), A (6.8%), Z (3.4%), M9 (2.5%), and M8 (1.7%).[97]
Korean people
[edit]Korean populations in modern Korea can be traced to many origins from the people of the Mumun period to the Yemaek people.[98] In modern times, Koreans are related to other populations found in Northeast Asia, however according to recent studies, ancient Koreans included populations related to the Yayoi people,[99] Jōmon people,[100] Siberian influx[101] etc.

Autosomal DNA
[edit]Generally speaking, modern Koreans' genetic ancestry is mostly dominated by Northeast Asian DNA with a small mix of Southern Jōmon-like ancestry (6% ±3%).[102][103] Evidence for both Northern and Southern mtDNA and Y-DNA haplogroups has been observed in Koreans, similar to the Japanese,[67] with the latter also being the closest group to the Koreans in the approximate region due to the overlap of Northeastern DNA and the presence of Jōmon-like M7a haplogroup [ja].[89] It is believed that the Jōmon-like ancestry was prominent during Neolithic period of Korea[104] with percentage as high as 34% (±7%),[102] but diminished over time due to incoming populations from the north.
Ancient genome comparisons revealed that the genetic makeup of Koreans can be best described as an admixture between Northeast Asian hunter-gatherers and an influx of rice-farming Southeast Asian agriculturalists from the Yangtze river valley.[93][105] This is supported by archaeological, historical and linguistic evidence, which suggests that the direct ancestors of Koreans were proto-Koreans who inhabited the northeastern region of China (situated near the Liao River) and the northern part of the Korean peninsula during the Neolithic (8,000–1,000 BC) and Bronze (1,500–400 BC) Ages,[106] who later mixed with the Jōmon-like natives in the southern part of the peninsula before the Three Kingdoms period of Korea.[102]
Paternal lineages
[edit]Studies of polymorphisms in the human Y-chromosome have so far produced evidence to suggest that the Korean people have a long history as a distinct, mostly endogamous ethnic group, with successive waves of people moving to the peninsula and three major Y-chromosome haplogroups.[107] A majority of Koreans belong to subclades of haplogroup O-M175 (ca. 79% in total,[106][108] with about 42%[108] to 44%[106] belonging to haplogroup O2-M122, about 31%[106] to 32%[108] belonging to haplogroup O1b2-M176, and about 2%[106] to 3%[108] belonging to haplogroup O1a-M119), while a significant minority belong to subclades of haplogroup C2-M217 (ca. 12%[106] to 13%[108] in total). Other Y-DNA haplogroups, including haplogroup N-M231, haplogroup D-M55, and haplogroup Q-M242, are also found in smaller proportions of present-day Koreans.[109][110][111]
Maternal lineages
[edit]Studies of Korean mitochondrial DNA lineages have shown that there is a high frequency of Haplogroup D4, followed by haplogroup B, and then haplogroup A and haplogroup G. Haplogroups with lower frequency include N9, Y, F, D5, M7, M8, M9, M10, M11, R11, C, and Z.[112][113][114]
Mongolic peoples
[edit]The ethnogenesis of Mongolic peoples is largely linked with the expansion of Ancient Northeast Asians. They subsequently came into contact with other groups, notably Sinitic peoples to their South and Western Steppe Herders to their far West. The Mongolians pastoralist lifestyle, may in part be derived from the Western Steppe Herders, but without much geneflow between these two groups, suggesting cultural transmission.[115][116] The Mongols are believed to be the descendants of the Xianbei and the proto-Mongols. The former term includes the Mongols proper (also known as the Khalkha Mongols), Oirats, the Kalmyk people and the Southern Mongols. The latter comprises the Abaga Mongols, Abaganar, Aohans, Baarins, Gorlos Mongols, Jalaids, Jaruud, Khishigten, Khuuchid, Muumyangan and Onnigud. The Daur people are descendants of the para-Mongolic Khitan people.[117]
Paternal lineages
[edit]The majority of Mongols in Mongolia and Russia belong to subclades of haplogroup C-M217,[118] followed by lower frequency of O-M175 and N-M231.[119] A minority belongs to haplogroup Q-M242, and a variety of West Eurasian haplogroups.[120]
Maternal lineages
[edit]The maternal haplogroups are diverse but similar to other northern Asian populations, including Haplogroup D, Haplogroup C, Haplogroup B, and Haplogroup A, which are shared among indigenous American and Asian populations.[121] West Eurasian mtDNA haplogroups makes up a some minority percentages. Haplogroup HV, Haplogroup U, Haplogroup K, Haplogroup I, Haplogroup J are all found in Mongolic people.[122]
Han Chinese
[edit]
The origins of the Han Chinese primarily trace back to Neolithic Yellow River farmers, who descended from Ancient Northern East Asians (ANEA), and Neolithic groups near the Yangtze, who descended from Ancient Southern East Asians (ASEA).[124][125][126][127][128] Today's modern Han Chinese can be colloquially categorized into two subgroups, Northern and Southern Han Chinese, although it is a clinal population with no significant distinction.[129] The Han Chinese cluster retains a level of singularity with its admixture of ANEA and ASEA ancestries which is unique to the group[127] with Southern Han having dual ancestry from Northern Han and southern non-Han natives.[130] Compared to other East Asian populations, the Northern Han Chinese cluster is placed closer to the "Korean/Mainland Japanese" cluster in terms of a correlative genetic relationship (mostly due to the overlap of ANEA), but is also quite distinguishable from them genetically,[124][36] due to the presence of ASEA ancestry[131] and the absence of Jōmon ancestry. The Southern Han Chinese also share more alleles with Thai and other Kra–Dai peoples according to principal component analysis than Northern Han Chinese.[36]
The genetic makeup of the modern Han Chinese is not purely uniform in terms of physical appearance and biological structure due to the vast geographical expanse of China and the migratory percolations that have occurred throughout it over the last few millennia. This has also engendered the emergence and evolution of the diverse multiplicity of assorted Han subgroups found throughout the various regions of modern China today. Comparisons between the Y chromosome single-nucleotide polymorphisms (SNPs) and mitochondrial DNA (mtDNA) of modern Northern Han Chinese and 3000 year old Hengbei ancient samples from China's Central Plains show that they are extremely similar to each other. These findings demonstrate that the core fundamental structural basis that shaped the genetic makeup of the present-day Northern Han Chinese was already formed more than three thousand years ago.[129]
Studies of DNA remnants from the Central Plains area of China 3000 years ago show close affinity between that population and those of Northern Han today in both the Y-DNA and mtDNA. Both Northern and Southern Han show similar Y-DNA genetic structure.[132]
Northern Han Chinese populations also have some West Eurasian admixture,[133] especially Han Chinese populations in Shaanxi (~2%-4.6%)[134] and Liaoning (~2%).[135] During the Zhou dynasty, or earlier, peoples with paternal haplogroup Q-M120 also contributed to the ethnogenesis of Han Chinese people. This haplogroup is implied to be widespread in the Eurasian steppe and north Asia since it is found among Cimmerians in Moldova and Bronze Age natives of Khövsgöl. But it is currently near-absent in these regions except for East Asia. In modern China, haplogroup Q-M120 can be found in the northern and eastern regions.[136] [137] Other Y-DNA haplogroups that have been found with notable frequency in samples of Han Chinese include O-P203 (15/165 = 9.1%, 217/2091 = 10.38%,[138] 47/361 = 13.0%), C-M217 (10/168 = 6.0%, 27/361 = 7.5%, 176/2091 = 8.42%,[138] 187/1730 = 10.8%, 20/166 = 12.0%), N-M231 (6/166 = 3.6%, 94/2091 = 4.50%,[138] 18/361 = 5.0%, 117/1729 = 6.8%, 17/165 = 10.3%), O-M268(xM95, M176) (78/2091 = 3.73%,[138] 54/1147 = 4.7%,[139] 8/168 = 4.8%, 23/361 = 6.4%, 12/166 = 7.2%), and Q-M242 (2/168 = 1.2%, 49/1729 = 2.8%, 61/2091 = 2.92%,[138] 12/361 = 3.3%, 48/1147 = 4.2%[139]).
However, the mtDNA of Han Chinese increases in diversity as one looks from Northern to Southern China, which suggests that the influx of male Han Chinese migrants intermarried with the local female non-Han aborigines after arriving in what is now modern-day Guangdong, Fujian, and other regions of Southern China.[140][141] Despite this, tests comparing the genetic profiles of Northern Han, Southern Han, and non-Han southern natives determined that haplogroups O1b-M110, O2a1-M88 and O3d-M7, which are prevalent in non-Han southern natives, were only observed in some Southern Han Chinese (4% on average), but not in the Northern Han genetic profile. Therefore, this proves that the male contribution of the southern non-Han natives in the Southern Han genetic profile is limited, assuming that the frequency distribution of Y lineages in southern non-Han natives represents that prior to the expansion of Han culture two thousand years ago from the north.[140][142]
A recent, and to date the most extensive, genome-wide association study of the Han population, shows that geographic-genetic stratification from north to south has occurred and centrally placed populations act as the conduit for outlying ones.[143] Ultimately, with the exception in some ethnolinguistic branches of the Han Chinese, such as Pinghua and Tanka people,[144] there is a "coherent genetic structure" found in the entirety of the modern Han Chinese populace.[145] Although admixture proportions can vary according to geographic region, the average genetic distance between various Han Chinese populations is much lower than between European populations, for example.[146]
Autosomal DNA
[edit]A 2018 study calculated pairwise FST (a measure of genetic difference) based on genome-wide SNPs, among the Han Chinese (Northern Han from Beijing and Southern Han from Hunan, Jiangsu and Fujian provinces), Japanese and Korean populations sampled. It found that the smallest FST value was between Northern Han Chinese (Beijing) (CHB) and Southern Han (Hunan, Fujian, etc.) Chinese (CHS) (FST[CHB-CHS] = 0.0014), while CHB and Korean (KOR) (FST[CHB-KOR] = 0.0026) and between KOR and Japanese (JPT) (FST[JPT-KOR] = 0.0033). Generally, pairwise FST between Han Chinese, Japanese and Korean (0.0026~ 0.0090) are greater than that within Han Chinese (0.0014). These results suggested Han Chinese, Japanese and Korean are different in terms of genetic make-up, and the differences among the three groups are much larger than that between Northern and Southern Han Chinese.[147] Nonetheless, there is also genetic diversity among the Southern Han Chinese. The genetic composition of the Han population in Fujian might not accurately represent that of the Han population in Guangdong.

Another study shows that the Northern and Southern Han Chinese are genetically close to each other and it finds that the genetic characteristics of present-day Northern Han Chinese were already formed prior to three thousand years ago in the Central Plain area.[149]
A recent genetic study on the remains of people (~4,000 years BP) from the Mogou site in the Gansu-Qinghai (or Ganqing) region of China revealed more information on the genetic contributions of these ancient Di-Qiang people to the ancestors of the Northern Han. It was deduced that 3,300 to 3,800 years ago some Mogou people had merged into the ancestral Han population, resulting in the Mogou people being similar to some Northern Han in sharing up to ~33% paternal (O3a) and ~70% maternal (D, A, F, M10) haplogroups. The mixing ratio was possibly 13–18%.[150]
The estimated contribution of Northern Han to Southern Han is substantial in both paternal and maternal lineages and a geographic cline exists for mtDNA. As a result, the Northern Han are one of the primary contributors to the gene pool of the Southern Han. However, it is noteworthy that the expansion process was not only dominated by males, as is shown by both contribution of the Y-chromosome and the mtDNA from Northern Han to Southern Han. Northern Han Chinese and Southern Han Chinese exhibit both Ancient Northern East Asian and Ancient Southern East Asian ancestries.[151] These genetic observations are in line with historical records of continuous and large migratory waves of Northern China inhabitants escaping warfare and famine, to Southern China. Aside from these large migratory waves, other smaller southward migrations occurred during almost all periods in the past two millennia.[152] A study by the Chinese Academy of Sciences into the gene frequency data of Han subpopulations and ethnic minorities in China showed that Han subpopulations in different regions are also genetically quite close to the local ethnic minorities, suggesting that in many cases, ethnic minorities ancestry had mixed with Han, while at the same time, the Han ancestry had also mixed with the local ethnic minorities.[153]
Han Chinese, similar to other East Asian populations, have inherited West Eurasian ancestry, around 2.8% in Northern Han Chinese and around 1.7% in Southern Han Chinese, compared to the 2.2% West Eurasian ancestry in the Japanese and the 1.6% West Eurasian ancestry in the Korean people.[154]
An extensive, genome-wide association study of the Han population in 2008, shows that geographic-genetic stratification from north to south has occurred and centrally placed populations act as the conduit for outlying ones.[155] Ultimately, with the exception in some ethnolinguistic branches of the Han Chinese, such as Pinghua, there is "coherent genetic structure" (homogeneity) in all Han Chinese.[156]
Paternal lineages
[edit]The major haplogroups of Han Chinese belong to subclades of Haplogroup O-M175. Y-chromosome O2-M122 is a common DNA marker in Han Chinese, as it appeared in China in prehistoric times, and is found in approximately 50% of Chinese males, with frequencies tending to be high toward the east of the country, ranging from 29.7% to 52% in Han from Southern and Central China, to 55–68% in Han from the eastern and northeastern Chinese mainland and Taiwan.[157]
Other Y-DNA haplogroups that have been found with notable frequency in samples of Han Chinese include O-P203 (9.1–13.0%), C-M217 (6.0–12.0%), N-M231 (3.6–10.3%), O-M268(xM95, M176) (4.7–7%), and Q-M242 (2/168 = 1.2–4.2%).[139][157]
Maternal lineages
[edit]The mitochondrial-DNA haplogroups of the Han Chinese can be classified into the Northern East Asian-dominating haplogroups, including A, C, D, G, M8, M9, and Z, and the Southern East Asian-dominating haplogroups, including B, F, M7, N*, and R.[152]
These haplogroups account for 52.7% and 33.85% of those in the Northern Han, respectively. Haplogroup D is the modal mtDNA haplogroup among Northern East Asians. Among these haplogroups, D, B, F, and A were predominant in the Northern Han, with frequencies of 25.77%, 11.54%, 11.54%, and 8.08%, respectively.
However, in the Southern Han, the Northern and Southern East Asian-dominating mtDNA haplogroups accounted for 35.62% and 51.91%, respectively. The frequencies of haplogroups D, B, F, and A reached 15.68%, 20.85%, 16.29%, and 5.63%, respectively.[149][158][159][160][161]
Tibetan peoples
[edit]The ethnic roots of Tibetans can be traced back to a deep Eastern Asian lineage representing the indigenous population of the Tibetan plateau since c. 40,000 to 30,000 years ago, and arriving Neolithic farmers from the Yellow River within the last 10,000 years associated, and which can be associated with having introduced the Sino-Tibetan languages. Modern Tibetans derive up to 20% from Paleolithic Tibetans, with the remaining 80% being primarily derived from Yellow River farmers. The present-day Tibetan gene pool was formed at least 5,100 years BP.[162][163]
Paternal lineage
[edit]Tibetan males predominantly belong to the paternal lineage D-M174 followed by lower amounts of O-M175.[164]
Maternal lineage
[edit]Tibetan females belong mainly to the Northeast Asian maternal haplogroups M9a1a, M9a1b, D4g2, D4i and G2ac, showing continuity with ancient middle and upper Yellow River populations.[165]
Turkic peoples
[edit]
Linguistic and genetic evidence strongly suggests an early presence of Turkic peoples in eastern Mongolia.[166] The genetic evidence suggests that the Turkification of Central Asia was carried out by East Asian dominant minorities migrating out of Mongolia.[49]
Genetic data found that almost all modern Turkic-speaking peoples retained at least some shared ancestry associated with "Southern Siberian and Mongolian" (SSM) populations, supporting this region as the "Inner Asian Homeland (IAH) of the pioneer carriers of Turkic languages" which subsequently expanded into Central Asia.[167]
An Ancient Northeast Asian origin of the early Turkic peoples has been corroborated in multiple recent studies. Early and medieval Turkic groups however exhibited a wide range of both (Northern) East Asian and West Eurasian genetic origins, in part through long-term contact with neighboring peoples such as Iranian, Mongolic, Tocharian, Uralic and Yeniseian peoples, and others.[168][169][170][171][172][173]
Paternal lineages
[edit]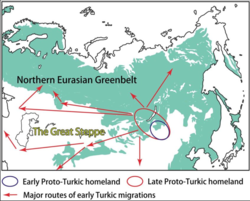
Common Y-DNA haplogroups in Turkic peoples are Haplogroup N-M231 (found with especially high frequency among Turkic peoples living in present-day Russia, especially among Siberian Tatars, as Zabolotnie Tatars have one of the highest frequencies of this haplogroup, second only to Samoyedic Nganasans ), Haplogroup C-M217 (especially in Central Asia, and in particular, Kazakhstan, also in Siberia among Siberian Tatars), Haplogroup Q-M242 (especially in Southern Siberia among the Siberian Tatars, also quite frequent among Lipka Tatars and among Turkmens and the Qangly tribe of Kazakhs), and Haplogroup O-M175 (especially among Turkic peoples living in present-day China, the Naiman tribe of Kazakhs and Siberian Tatars). Some groups also have Haplogroup R1b (notably frequent among the Teleuts, Siberian Tatars, and Kumandins of Southern Siberia, the Bashkirs of the Southern Ural region of Russia, and the Qypshaq tribe of Kazakhs), Haplogroup R1a (notably frequent among the Kyrgyz, Altaians, Siberian Tatars, Lipka Tatars, Volga Tatars, Crimean Tatars and several other Turkic peoples living in present-day Russia), Haplogroup J-M172 (especially frequent among Uyghurs, Azerbaijanis, and Turkish people), and Haplogroup D-M174 (especially among Yugurs, but also observed regularly with low frequency among Southern Altaians, Nogais, Kazakhs, and Uzbeks).[174][175]
Relationship to other Asia-Pacific and Native American populations
[edit]Central Asians
[edit]The oldest modern human genome found in Central Asia belongs to the deeply East Eurasian IUP-affiliated Ust'Ishim man. The population affiliated with this specimen is inferred to have not contributed to modern human populations. During the late Upper Paleolithic period, geneflow from Ancient North Eurasians (ANE) played a significant role in the genetic makeup of Central Asia. The ANE carried both Upper Paleolithic European and East/Southeast Asian ancestry. Post-Paleolithic geneflow included movements of Paleo-Siberian and Northeast Asian groups into Central Asia, admixing with local ANE-rich groups resulting in the formation of the Botai genetic grouping, with close affinities to West Siberian hunter-gatherers (WSHG). Subsequently, massive geneflow from Western Steppe Herders from eastern and central Europe is associated with the formation of the Andronovo culture and the spread of Indo-Iranian languages. Interactions between pre-existing groups with Andronovo and Paleo-Siberian tribes, resulted in the origin of early Scythians.[176] During the Iron Age, the Turkification of Central Asia was carried out by East Asian dominant minorities migrating out of Mongolia.[177] The Turkic-speaking Central Asian populations, such as Kyrgyz, Kazakhs, Uzbeks, and Turkmens share more of their gene pool with various East Asian and Siberian populations than with West Asian or European populations.[178]
Native Americans
[edit]All Native American populations descend from an ancient Paleo-Siberian group which emerged by the merger of Ancient East Asians and Ancient North Eurasians. While the East Asian-like ancestry is best represented by Ancient Northern East Asians (Amur14k), it may also include ancestry from further South prior to the divergence between Southern and Northern East Asians. Beyond that, there may be a small and variable amount of deeply branching East Asian admixture best represented by the Onge, Papuans or the Tianyuan man. This deep ghost component has been dubbed as "population Y".[67][179][180][181][182][183][1]
South Asians
[edit]The genetic makeup of modern South Asians can be described as a combination of West Eurasian ancestries with divergent East Eurasian ancestries. The latter primarily include an indigenous South Asian component (termed Ancient Ancestral South Indians, short "AASI") that is distantly related to the Andamanese peoples, as well as to East Asians and Aboriginal Australians, and further include additional, regionally variable East/Southeast Asian components.[184][7] The East Asian-related ancestry component forms the major ancestry among Tibeto-Burmese and Khasi-Aslian speakers in the Himalayan foothills and Northeast India,[185][186] and is generally distributed throughout South Asia at lower frequency, with substantial presence in Mundari-speaking groups.[185][186][187] Southern East Asian ancestry is primarily associated with the dispersal of Austroasiatic rice farmers, which migrated from Southeast Asia into India.[188] Multiple researches indicate that the Austroasiatic populations in India are derived from (mostly male dominated) migrations from Southeast Asia during the Holocene.[189][188][190][191][192][193] According to Van Driem (2007), "...the mitochondrial picture indicates that the Munda maternal lineage derives from the earliest human settlers on the Subcontinent, whilst the predominant Y chromosome haplogroup argues for a Southeast Asian paternal homeland for Austroasiatic language communities in India."[190]: 7
Geneflow from Southeast Asians (particularly Austroasiatic groups) to South Asian peoples is associated with the introduction of rice-agriculture to South Asia. There is significant cultural, linguistic, and political Austroasiatic influence on early India, which can also be observed by the presence of Austroasiatic loanwords within Indo-Aryan languages.[194][195]
Southeast Asians
[edit]
Southeast Asians represent one of the most closely related groups to East Asians, with both being referred to as East/Southeast Asian. While East Asians primarily derive from both ANEA and ASEA components, Southeast Asians derive most of their ancestry from the ASEA component with variable amounts of deeper branching East Asian-like admixture (mostly Onge/Hoabinhian-like). While Hoabinhian-like ancestry is associated with indigenous hunter-gatherers, ASEA ancestry spread mostly with Neolithic expansions associated with Austroasiatic and Austronesian groups.[41][197][17][1]
Evidence for more complex Mesolithic migration patterns are evident in the remains of a hunter-gatherer specimen from Maritime Southeast Asia, South Sulawesi, which was found to have ancestry from two deeply diverged East Eurasian lineages. The remains had approximately c. 50% "Tianyuan/Onge" ancestry and c. 50% Papuan-like ancestry.[198]
There is also evidence for low proportions (~5%) of South Asian-associated "SAS ancestry" (best examplified by modern South Indian groups such as Irula or Mala) among specific Mainland Southeast Asian (MESA) ethnic groups (~2–16% as inferred by qpAdm), likely as a result of cultural diffusion; mainly through maritime trade and Indianized kingdoms of Southeast Asia. Overall, the geneflow event is estimated to have happened between 500 and 1000 YBP.[199]

Australasians
[edit]Melanesians and Aboriginal Australians are deeply related to East Asians. Genetic studies have revealed that Australasians descended from the same Eastern Eurasian source population as East Asians and indigenous South Asians (AASI).[7] The 'Australasian', 'Ancient Ancestral South Indian', and 'East and Southeast Asian' lineages display a closer genetic relationship to each other than to any non-Asian lineages, as well as being closer to each other than to any of the early East Eurasian IUP lineages (Bacho Kiro etc.), and together represent the main branches of "Asian-related ancestry", which diverged from each other >40,000 years ago.[200]
References
[edit]- ^ a b c d e f g h i j k l Bennett, Andrew E.; Liu, Yichen; Fu, Qiaomei (4 December 2024). "Reconstructing the Human Population History of East Asia through Ancient Genomics". Elements in Ancient East Asia. doi:10.1017/9781009246675. ISBN 978-1-009-24667-5.
- ^ Wang CC, Li H (June 2013). "Inferring human history in East Asia from Y chromosomes". Investigative Genetics. 4 (1): 11. doi:10.1186/2041-2223-4-11. PMC 3687582. PMID 23731529.
- ^ Di D, Sanchez-Mazas A, Currat M (November 2015). "Computer simulation of human leukocyte antigen genes supports two main routes of colonization by human populations in East Asia". BMC Evolutionary Biology. 15 (1): 240. Bibcode:2015BMCEE..15..240D. doi:10.1186/s12862-015-0512-0. PMC 4632674. PMID 26530905.
- ^ a b c d e Vallini L, Marciani G, Aneli S, Bortolini E, Benazzi S, Pievani T, Pagani L (April 2022). "Genetics and Material Culture Support Repeated Expansions into Paleolithic Eurasia from a Population Hub Out of Africa". Genome Biology and Evolution. 14 (4). doi:10.1093/gbe/evac045. PMC 9021735. PMID 35445261.
- ^ Yang, Melinda A.; Gao, Xing; Theunert, Christoph; Tong, Haowen; Aximu-Petri, Ayinuer; Nickel, Birgit; Slatkin, Montgomery; Meyer, Matthias; Pääbo, Svante; Kelso, Janet; Fu, Qiaomei (23 October 2017). "40,000-Year-Old Individual from Asia Provides Insight into Early Population Structure in Eurasia". Current Biology. 27 (20): 3202–3208.e9. doi:10.1016/j.cub.2017.09.030. ISSN 0960-9822. PMC 6592271.
This is consistent with a split time of 40,000–80,000 years ago estimated for European and Asian populations based on mutation rates estimated from de novo mutations [13, 14, 15].
- ^ a b Vallini, Leonardo; Zampieri, Carlo; Shoaee, Mohamed Javad; Bortolini, Eugenio; Marciani, Giulia; Aneli, Serena; Pievani, Telmo; Benazzi, Stefano; Barausse, Alberto; Mezzavilla, Massimo; Petraglia, Michael D.; Pagani, Luca (25 March 2024). "The Persian plateau served as hub for Homo sapiens after the main out of Africa dispersal". Nature Communications. 15 (1): 1882. Bibcode:2024NatCo..15.1882V. doi:10.1038/s41467-024-46161-7. ISSN 2041-1723. PMC 10963722. PMID 38528002.
We simulate the OOA 60 kya, with Basal Eurasians (BEA in Supplementary Fig. 4) splitting soon after (57.5 kya) and the split between EEC and WEC, with the former leaving the Hub18, 46 kya (allowing the time for them to reach Ust'Ishim and Bacho Kiro by ~45 kya).
- ^ a b c d e Yang MA (6 January 2022). "A genetic history of migration, diversification, and admixture in Asia". Human Population Genetics and Genomics. 2 (1): 1–32. doi:10.47248/hpgg2202010001. ISSN 2770-5005.
...In contrast, mainland East and Southeast Asians and other Pacific islanders (e.g., Austronesian speakers) are closely related to each other [9,15,16] and here denoted as belonging to an East and Southeast Asian (ESEA) lineage (Box 2). …the ESEA lineage differentiated into at least three distinct ancestries: Tianyuan ancestry which can be found 40,000–33,000 years ago in northern East Asia, ancestry found today across present-day populations of East Asia, Southeast Asia, and Siberia, but whose origins are unknown, and Hòabìnhian ancestry found 8,000–4,000 years ago in Southeast Asia, but whose origins in the Upper Paleolithic are unknown.
- ^ a b c d Aoki K, Takahata N, Oota H, Wakano JY, Feldman MW (August 2023). "Infectious diseases may have arrested the southward advance of microblades in Upper Palaeolithic East Asia". Proceedings. Biological Sciences. 290 (2005): 20231262. doi:10.1098/rspb.2023.1262. PMC 10465978. PMID 37644833.
A single major migration of modern humans into the continents of Asia and Sahul was strongly supported by earlier studies using mitochondrial DNA, the non-recombining portion of Y chromosomes, and autosomal SNP data [42–45]. Ancestral Ancient South Indians with no West Eurasian relatedness, East Asians, Onge (Andamanese hunter–gatherers) and Papuans all derive in a short evolutionary time from the eastward dispersal of an out-of-Africa population [46,47], although Europeans and East Asians are suggested to share more recent common ancestors than with Papuans [48]. The HUGO (Human Genome Organization) Pan-Asian SNP consortium [44] investigated haplotype diversity within present-day Asian populations and found a strong correlation with latitude, with diversity decreasing from south to north. The correlation continues to hold when only mainland Southeast Asian and East Asian populations are considered, and is perhaps attributable to a serial founder effect [50]. These observations are consistent with the view that soon after the single eastward migration of modern humans, East Asians diverged in southern East Asia and dispersed northward across the continent.
- ^ Genetics and material culture support repeated expansions into Paleolithic Eurasia from a population hub out of Africa, Vallini et al. 2022 (4 April 2022) Quote: "Taken together with a lower bound of the final settlement of Sahul at 37 ka (the date of the deepest population splits estimated by Malaspinas et al. 2016), it is reasonable to describe Papuans as either an almost even mixture between East Asians and a lineage basal to West and East Asians occurred sometimes between 45 and 38 ka, or as a sister lineage of East Asians with or without a minor basal OoA or xOoA contribution. We here chose to parsimoniously describe Papuans as a simple sister group of Tianyuan, cautioning that this may be just one out of six equifinal possibilities."
- ^ "Almost all living people outside of Africa trace back to a single migration more than 50,000 years ago". science.org. Retrieved 19 August 2022.
- ^ Metspalu M, Kivisild T, Metspalu E, Parik J, Hudjashov G, Kaldma K, et al. (August 2004). "Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans". BMC Genetics. 5 (1): 26. doi:10.1186/1471-2156-5-26. PMC 516768. PMID 15339343.
- ^ a b Osada N, Kawai Y (2021). "Exploring models of human migration to the Japanese archipelago using genome-wide genetic data". Anthropological Science. 129 (1): 45–58. doi:10.1537/ase.201215. S2CID 234247309.
Via the southern route, ancestors of current Asian populations reached Southeast Asia and a part of Oceania around 70000–50000 years ago, probably through a coastal dispersal route (Bae et al., 2017). The oldest samples providing the genetic evidence of the northern migration route come from a high-coverage genome sequence of individuals excavated from the Yana RHS site in northeastern Siberia (Figure 2), which is about 31600 years old (Sikora et al., 2019). A wide range of artifacts, including bone crafts of wooly rhinoceros and mammoths, were excavated at the site (Pitulko et al., 2004). The analysis of genome sequences showed that the samples were deeply diverged from most present-day East Asians and more closely related to present-day Europeans, suggesting that the population reached the area through a route different from the southern route. A 24000-year-old individual excavated near Lake Baikal (Figure 2), also known as the Mal'ta boy, and 17000-year-old individuals from the Afontova Gora II site (Afontova Gora 2 and 3) showed similar genetic features to the Yana individuals (Raghavan et al., 2014; Fu et al., 2016; Sikora et al., 2019). Interestingly, genetic data suggested that Yana individuals received a large amount of gene flow from the East Asian lineage (Sikora et al., 2019; Yang et al., 2020).
- ^ Gakuhari T, Nakagome S, Rasmussen S, Allentoft ME, Sato T, Korneliussen T, et al. (August 2020). "Ancient Jomon genome sequence analysis sheds light on migration patterns of early East Asian populations". Communications Biology. 3 (1): 437. doi:10.1038/s42003-020-01162-2. PMC 7447786. PMID 32843717.
Population genomic studies on present-day humans7,8 have exclusively supported the southern route origin of East Asian populations.
- ^ Larena M, Sanchez-Quinto F, Sjödin P, McKenna J, Ebeo C, Reyes R, et al. (March 2021). "Multiple migrations to the Philippines during the last 50,000 years". Proceedings of the National Academy of Sciences of the United States of America. 118 (13): e2026132118. Bibcode:2021PNAS..11826132L. doi:10.1073/pnas.2026132118. PMC 8020671. PMID 33753512.
- ^ Bae CJ, Douka K, Petraglia MD (December 2017). "On the origin of modern humans: Asian perspectives". Science. 358 (6368): eaai9067. doi:10.1126/science.aai9067. PMID 29217544. S2CID 4436271.
- ^ Demeter, Fabrice; Shackelford, Laura L.; Bacon, Anne-Marie; Duringer, Philippe; Westaway, Kira; Sayavongkhamdy, Thongsa; Braga, José; Sichanthongtip, Phonephanh; Khamdalavong, Phimmasaeng; Ponche, Jean-Luc; Wang, Hong; Lundstrom, Craig; Patole-Edoumba, Elise; Karpoff, Anne-Marie (4 September 2012). "Anatomically modern human in Southeast Asia (Laos) by 46 ka". Proceedings of the National Academy of Sciences. 109 (36): 14375–14380. Bibcode:2012PNAS..10914375D. doi:10.1073/pnas.1208104109. ISSN 0027-8424. PMC 3437904. PMID 22908291.
Inferences from nuclear (51), Y chromosome (52), and mitochondrial genome (53) data support an early migration of modern humans out of Africa and into Southeast Asia using a southern route by at least 60 ka. Patterns of genetic variation in recent human populations (11, 54, 55) recognize Southeast Asia as an important source for the peopling of East Asia and Australasia via a rapid, early settlement.
- ^ a b c d e f Yang, Melinda A. (6 January 2022). "A genetic history of migration, diversification, and admixture in Asia". Human Population Genetics and Genomics. 2 (1). doi:10.47248/hpgg2202010001. ISSN 2770-5005.
- ^ a b Gakuhari, Takashi; Nakagome, Shigeki; Rasmussen, Simon; Allentoft, Morten E.; Sato, Takehiro; Korneliussen, Thorfinn; Chuinneagáin, Blánaid Ní; Matsumae, Hiromi; Koganebuchi, Kae; Schmidt, Ryan; Mizushima, Souichiro; Kondo, Osamu; Shigehara, Nobuo; Yoneda, Minoru; Kimura, Ryosuke (25 August 2020). "Ancient Jomon genome sequence analysis sheds light on migration patterns of early East Asian populations". Communications Biology. 3 (1): 1–10. doi:10.1038/s42003-020-01162-2. ISSN 2399-3642. PMC 7447786. PMID 32843717.
Population genomic studies on present-day humans7,8 have exclusively supported the southern route origin of East Asian populations.
- ^ a b Watanabe, Yusuke; Ohashi, Jun (17 March 2023). "Modern Japanese ancestry-derived variants reveal the formation process of the current Japanese regional gradations". iScience. 26 (3). doi:10.1016/j.isci.2023.106130. ISSN 2589-0042. PMC 9984562. PMID 36879818.
Whole-genome analyses extracted from the remains of the Jomon people showed that they were highly differentiated from other East Asians, forming a basal lineage to East and Northeast Asians.8,10,11 The genetic relationship between Jomon individuals and other East Asians suggests that the ancestral population of the Jomon people is one of the earliest wave migrants who might have taken a coastal route from Southeast Asia toward East Asia.
- ^ a b Matsumura, Hirofumi; Hung, Hsiao-chun; Higham, Charles; Zhang, Chi; Yamagata, Mariko; Nguyen, Lan Cuong; Li, Zhen; Fan, Xue-chun; Simanjuntak, Truman; Oktaviana, Adhi Agus; He, Jia-ning; Chen, Chung-yu; Pan, Chien-kuo; He, Gang; Sun, Guo-ping (5 February 2019). "Craniometrics Reveal "Two Layers" of Prehistoric Human Dispersal in Eastern Eurasia". Scientific Reports. 9 (1): 1451. doi:10.1038/s41598-018-35426-z. hdl:1885/156483. ISSN 2045-2322.
...ancient people perhaps of the "first layer" with Australo-Papuan features moved into Siberia and subsequently adapted to the extremely cold climate during the Last Glacial Maximum (LGM) of 24– 16 kya.
- ^ a b Wang, Tianyi; Wang, Wei; Xie, Guangmao; Li, Zhen; Fan, Xuechun; Yang, Qingping; Wu, Xichao; Cao, Peng; Liu, Yichen; Yang, Ruowei; Liu, Feng; Dai, Qingyan; Feng, Xiaotian; Wu, Xiaohong; Qin, Ling (8 July 2021). "Human population history at the crossroads of East and Southeast Asia since 11,000 years ago". Cell. 184 (14): 3829–3841.e21. doi:10.1016/j.cell.2021.05.018. ISSN 0092-8674. PMID 34171307.
However, genetic sampling in Japan and southern China of populations associated craniometrically with the first layer show that they are more closely related genetically to second-layer East Asian populations, indicating that the two-layer model is not sufficient to describe the population movement, replacement, and mixture in prehistoric Asia.
- ^ a b Xu D, Li H (2017). Languages and Genes in Northwestern China and Adjacent Regions. Springer. p. 27. ISBN 978-981-10-4169-3. "In the study of Zhong et al. haplogroups O-M175, C-M130, D-M174 and N-M231 still suggests the substantial contribution of the southern route. However, the Central Asia and West Eurasia related haplogroups, such as haplogroups R-M207 and Q-M242, occur primarily in northwestern East Asia and their frequencies gradually decrease from west to east. In addition, the Y-STR diversities of haplogroups R-M207 and Q-M242 also indicate the existence of northern route migration about 18,000 years ago from Central Asia to North Asia, and recent population admixture along the Silk Road since about 3000 years ago (Piazza 1998)."
- ^ a b Wang, Chuan-Chao; Yeh, Hui-Yuan; Popov, Alexander N; Zhang, Hu-Qin; Matsumura, Hirofumi; Sirak, Kendra; Cheronet, Olivia; Kovalev, Alexey; Rohland, Nadin; Kim, Alexander M; Mallick, Swapan; Bernardos, Rebecca; Tumen, Dashtseveg; Zhao, Jing; Liu, Yi-Chang (22 February 2021). "Genomic Insights into the Formation of Human Populations in East Asia". Nature. 591 (7850). doi:10.1038/s41586-021-03336-2. Archived from the original on 7 March 2025.
- ^ a b c McColl, Hugh; Racimo, Fernando; Vinner, Lasse; Demeter, Fabrice; Gakuhari, Takashi; et al. (2018). "The prehistoric peopling of Southeast Asia". Science. 361 (6397). American Association for the Advancement of Science (AAAS): 88–92. Bibcode:2018Sci...361...88M. doi:10.1126/science.aat3628. hdl:10072/383365. ISSN 0036-8075. PMID 29976827. S2CID 206667111.
- ^ Lazaridis, Iosif; Belfer-Cohen, Anna; Mallick, Swapan; Patterson, Nick; Cheronet, Olivia; Rohland, Nadin; Bar-Oz, Guy; Bar-Yosef, Ofer; Jakeli, Nino (21 September 2018), Paleolithic DNA from the Caucasus reveals core of West Eurasian ancestry, bioRxiv, doi:10.1101/423079, retrieved 17 April 2025
- ^ Wang, Hongru; Yang, Melinda A.; Wangdue, Shargan; Lu, Hongliang; Chen, Honghai; Li, Linhui; Dong, Guanghui; Tsring, Tinley; Yuan, Haibing; He, Wei; Ding, Manyu; Wu, Xiaohong; Li, Shuai; Tashi, Norbu; Yang, Tsho (17 March 2023). "Human genetic history on the Tibetan Plateau in the past 5100 years". Science Advances. 9 (11): eadd5582. doi:10.1126/sciadv.add5582. PMC 10022901. PMID 36930720.
- ^ Hajdinjak, Mateja; Mafessoni, Fabrizio; Skov, Laurits; Vernot, Benjamin; Hübner, Alexander; Fu, Qiaomei; Essel, Elena; Nagel, Sarah; Nickel, Birgit; Richter, Julia; Moldovan, Oana Teodora; Constantin, Silviu; Endarova, Elena; Zahariev, Nikolay; Spasov, Rosen (7 April 2021). "Initial Upper Palaeolithic humans in Europe had recent Neanderthal ancestry". Nature. 592 (7853): 253–257. doi:10.1038/s41586-021-03335-3. hdl:11585/827583. ISSN 1476-4687.
- ^ Massilani, Diyendo; Skov, Laurits; Hajdinjak, Mateja; Gunchinsuren, Byambaa; Tseveendorj, Damdinsuren; Yi, Seonbok; Lee, Jungeun; Nagel, Sarah; Nickel, Birgit; Devièse, Thibaut; Higham, Tom; Meyer, Matthias; Kelso, Janet; Peter, Benjamin M.; Pääbo, Svante (30 October 2020). "Denisovan ancestry and population history of early East Asians". Science. 370 (6516): 579–583. doi:10.1126/science.abc1166.
- ^ Allentoft, Morten E.; Sikora, Martin; Refoyo-Martínez, Alba; Irving-Pease, Evan K.; Fischer, Anders; Barrie, William; Ingason, Andrés; Stenderup, Jesper; Sjögren, Karl-Göran; Pearson, Alice; Sousa da Mota, Bárbara; Schulz Paulsson, Bettina; Halgren, Alma; Macleod, Ruairidh; Jørkov, Marie Louise Schjellerup (10 January 2024). "Population genomics of post-glacial western Eurasia". Nature. 625 (7994): 301–311. doi:10.1038/s41586-023-06865-0. ISSN 1476-4687. PMC 10781627. PMID 38200295.
- ^ Yang, Melinda A.; Gao, Xing; Theunert, Christoph; Tong, Haowen; Aximu-Petri, Ayinuer; Nickel, Birgit; Slatkin, Montgomery; Meyer, Matthias; Pääbo, Svante; Kelso, Janet; Fu, Qiaomei (23 October 2017). "40,000-Year-Old Individual from Asia Provides Insight into Early Population Structure in Eurasia". Current Biology. 27 (20): 3202–3208.e9. doi:10.1016/j.cub.2017.09.030. ISSN 0960-9822. PMC 6592271. PMID 29033327.
- ^ Zhang X, Ji X, Li C, Yang T, Huang J, Zhao Y, et al. (July 2022). "A Late Pleistocene human genome from Southwest China". Current Biology. 32 (14): 3095–3109.e5. Bibcode:2022CBio...32E3095Z. doi:10.1016/j.cub.2022.06.016. PMID 35839766. S2CID 250502011. "In addition to the earliest southern settlement of AMHs in East Asia, ancient migration (40–18 kya) into East Asia via the “Northern Route” from West Eurasia was previously proposed. The “Northern Route” hypothesis would also explain where the subtle shared ancient north Eurasian (ANE) ancestry came from that is then also shared with Native Americans."
- ^ Villalba-Mouco, Vanessa; van de Loosdrecht, Marieke S.; Rohrlach, Adam B.; Fewlass, Helen; Talamo, Sahra; Yu, He; Aron, Franziska; Lalueza-Fox, Carles; Cabello, Lidia; Cantalejo Duarte, Pedro; Ramos-Muñoz, José; Posth, Cosimo; Krause, Johannes; Weniger, Gerd-Christian; Haak, Wolfgang (1 April 2023). "A 23,000-year-old southern Iberian individual links human groups that lived in Western Europe before and after the Last Glacial Maximum". Nature Ecology & Evolution. 7 (4): 597–609. doi:10.1038/s41559-023-01987-0. ISSN 2397-334X. PMC 10089921. PMID 36859553.
- ^ Huang, Xin (2021). "Dissecting dynamics and differences of selective pressures in the evolution of human pigmentation". Biology Open. 10 (2). doi:10.1242/bio.056523. PMC 7888712. PMID 33495209.
- ^ Liu, Jiuming; Bitsue, Habtom K.; Yang, Zhaohui (2024). "Skin colour: A window into human phenotypic evolution and environmental adaptation". Molecular Ecology. 33 (12): e17369. doi:10.1111/mec.17369. ISSN 1365-294X.
Positive selection for rs1800414-G in East Asia is thought to date back to the Late Palaeolithic period (~25–30 kya), following the northward migration of modern humans from South Asia (Yang et al., 2016).
- ^ a b Mao X, Zhang H, Qiao S, Liu Y, Chang F, Xie P, et al. (June 2021). "The deep population history of northern East Asia from the Late Pleistocene to the Holocene". Cell. 184 (12): 3256–3266.e13. doi:10.1016/j.cell.2021.04.040. PMID 34048699. S2CID 235226413.
- ^ a b c Huang, Xiufeng; Xia, Zi-Yang; Bin, Xiaoyun; He, Guanglin; Guo, Jianxin; Lin, Chaowen; Yin, Lianfei; Zhao, Jing; Ma, Zhuofei (8 November 2020), Genomic Insights into the Demographic History of Southern Chinese, doi:10.1101/2020.11.08.373225
- ^ Zhang, Ming; Fu, Qiaomei (1 June 2020). "Human evolutionary history in Eastern Eurasia using insights from ancient DNA". Current Opinion in Genetics & Development. Genetics of Human Origin. 62: 78–84. doi:10.1016/j.gde.2020.06.009. ISSN 0959-437X. PMID 32688244. S2CID 220671047.
- ^ He, Guang-Lin; Wang, Meng-Ge; Zou, Xing; Yeh, Hui-Yuan; Liu, Chang-Hui; Liu, Chao; Chen, Gang; Wang, Chuan-Chao (2023). "Extensive ethnolinguistic diversity at the crossroads of North China and South Siberia reflects multiple sources of genetic diversity". Journal of Systematics and Evolution. 61 (1): 230–250. doi:10.1111/jse.12827. ISSN 1674-4918.
- ^ Wong EH, Khrunin A, Nichols L, Pushkarev D, Khokhrin D, Verbenko D, et al. (January 2017). "Reconstructing genetic history of Siberian and Northeastern European populations". Genome Research. 27 (1): 1–14. doi:10.1101/gr.202945.115. PMC 5204334. PMID 27965293.
- ^ Sikora, Martin; Pitulko, Vladimir V.; Sousa, Vitor C.; Allentoft, Morten E.; Vinner, Lasse; Rasmussen, Simon; Margaryan, Ashot; Damgaard, Peter de Barros; Castro, Constanza de la Fuente (22 October 2018), The population history of northeastern Siberia since the Pleistocene, doi:10.1101/448829, hdl:1887/3198847, retrieved 14 November 2024
- ^ a b Tagore D, Aghakhanian F, Naidu R, Phipps ME, Basu A (March 2021). "Insights into the demographic history of Asia from common ancestry and admixture in the genomic landscape of present-day Austroasiatic speakers". BMC Biology. 19 (1): 61. doi:10.1186/s12915-021-00981-x. PMC 8008685. PMID 33781248.
- ^ "Direct genetic evidence of founding population reveals story of first Native Americans". University of Cambridge. 3 January 2018. Retrieved 9 June 2021.
- ^ Liu D, Duong NT, Ton ND, Van Phong N, Pakendorf B, Van Hai N, Stoneking M (September 2020). "Extensive Ethnolinguistic Diversity in Vietnam Reflects Multiple Sources of Genetic Diversity". Molecular Biology and Evolution. 37 (9): 2503–2519. doi:10.1093/molbev/msaa099. PMC 7475039. PMID 32344428.
- ^ Liu, Dang; Duong, Nguyen Thuy; Ton, Nguyen Dang; Phong, Nguyen Van; Pakendorf, Brigitte; Hai, Nong Van; Stoneking, Mark (28 November 2019), Extensive ethnolinguistic diversity in Vietnam reflects multiple sources of genetic diversity, doi:10.1101/857367, hdl:21.11116/0000-0006-4AD8-4, retrieved 14 November 2024
- ^ "Xiongnu People". britannica.com. Encyclopædia Britannica. Archived from the original on 11 March 2020. Retrieved 25 July 2015.
- ^ di Cosmo 2004: 186
- ^ a b c Savelyev, Alexander; Jeong, Choongwoon (7 May 2020). "Early nomads of the Eastern Steppe and their tentative connections in the West". Evolutionary Human Sciences. 2 (E20). doi:10.1017/ehs.2020.18. hdl:21.11116/0000-0007-772B-4. PMC 7612788. PMID 35663512. S2CID 218935871.
 Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. "Such a distribution of Xiongnu words may be an indication that both Turkic and Eastern Iranian-speaking groups were present among the Xiongnu in the earlier period of their history. Etymological analysis shows that some crucial components in the Xiongnu political, economic and cultural package, including dairy pastoralism and elements of state organization, may have been imported by the Eastern Iranians. Arguably, these Iranian-speaking groups were assimilated over time by the predominant Turkic-speaking part of the Xiongnu population. ... The genetic profile of published Xiongnu individuals speaks against the Yeniseian hypothesis, assuming that modern Yeniseian speakers (i.e. Kets) are representative of the ancestry components in the historical Yeniseian speaking groups in southern Siberia. In contrast to the Iron Age populations listed in Table 2, Kets do not have the Iranian-related ancestry component but harbour a strong genetic affinity with Samoyedic-speaking neighbours, such as Selkups (Jeong et al., 2018, 2019)."
Text was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. "Such a distribution of Xiongnu words may be an indication that both Turkic and Eastern Iranian-speaking groups were present among the Xiongnu in the earlier period of their history. Etymological analysis shows that some crucial components in the Xiongnu political, economic and cultural package, including dairy pastoralism and elements of state organization, may have been imported by the Eastern Iranians. Arguably, these Iranian-speaking groups were assimilated over time by the predominant Turkic-speaking part of the Xiongnu population. ... The genetic profile of published Xiongnu individuals speaks against the Yeniseian hypothesis, assuming that modern Yeniseian speakers (i.e. Kets) are representative of the ancestry components in the historical Yeniseian speaking groups in southern Siberia. In contrast to the Iron Age populations listed in Table 2, Kets do not have the Iranian-related ancestry component but harbour a strong genetic affinity with Samoyedic-speaking neighbours, such as Selkups (Jeong et al., 2018, 2019)."
- ^ Savelyev & Jeong 2020:"Specifically, individuals from Iron Age steppe and Xiongnu have an ancestry related to present-day and ancient Iranian/Caucasus/Turan populations in addition to the ancestry components derived from the Late Bronze Age populations. We estimate that they derive between 5 and 25% of their ancestry from this new source, with 18% for Xiongnu (Table 2). We speculate that the introduction of this new western Eurasian ancestry may be linked to the Iranian elements in the Xiongnu linguistic material, while the Turkic-related component may be brought by their eastern Eurasian genetic substratum." Table 2: Sintashta_MLBA, 0.239; Khovsgol LBA, 0.582; Gonur1 BA 0.178
- ^ a b Damgaard PB, Marchi N, Rasmussen S, Peyrot M, Renaud G, Korneliussen T, et al. (May 2018). "137 ancient human genomes from across the Eurasian steppes". Nature. 557 (7705): 369–374. Bibcode:2018Natur.557..369D. doi:10.1038/s41586-018-0094-2. hdl:1887/3202709. PMID 29743675. S2CID 13670282.
- ^ Lee, Juhyeon; Miller, Bryan K.; Bayarsaikhan, Jamsranjav; Johannesson, Erik; Miller, Alicia Ventresca; Warinner, Christina; Jeong, Choongwon (April 2023). "Genetic population structure of the Xiongnu Empire at imperial and local scales". Science Advances. 9 (15): eadf3904. Bibcode:2023SciA....9F3904L. doi:10.1126/sciadv.adf3904. PMC 10104459. PMID 37058560.
- ^ Rogers & Kaestle 2022.
- ^ Li H (2012). Y-Chromosome Genetic Diversity of the Ancient North Chinese populations (Report). China: Jilin University.
- ^ Lee J, Miller BK, Bayarsaikhan J, Johannesson E, Ventresca Miller A, Warinner C, Jeong C (April 2023). "Genetic population structure of the Xiongnu Empire at imperial and local scales". Science Advances. 9 (15). American Association for the Advancement of Science (AAAS): eadf3904. Bibcode:2023SciA....9F3904L. doi:10.1126/sciadv.adf3904. PMC 10104459. PMID 37058560.
- ^ Lee 2023: Haplogroup information for DA39 is located in Supplementary Materials document Data S1 (abk1900_Data_S1.xlsx), row 58. "Before this study, only one other individual from an elite square tomb had been analyzed in a genome-wide manner: DA39 from Tomb 1 at the imperial elite site of Gol Mod 2 in central-north Mongolia (13). This adult male, buried in one of the largest square tomb complexes excavated to date, surrounded by at least 27 satellite burials, and containing rare exotic items such as Roman glass bowls, was likely a chanyu, or ruler of the empire (73, 74)."
- ^ Rogers LL, Kaestle FA (2022). "Analysis of mitochondrial DNA haplogroup frequencies in the population of the slab burial mortuary culture of Mongolia (ca. 1100–300 BCE )". American Journal of Biological Anthropology. 177 (4): 644–657. doi:10.1002/ajpa.24478. ISSN 2692-7691. S2CID 246508594. " The first pattern is that the slab burial mtDNA frequencies are extremely similar to those of the aggregated Xiongnu populations and relatively similar to those of the various Bronze Age Mongolian populations, strongly supporting a population continuity hypothesis for the region over these time periods (Honeychurch, 2013)"
- ^ Keyser-Tracqui C, Crubézy E, Ludes B (August 2003). "Nuclear and mitochondrial DNA analysis of a 2,000-year-old necropolis in the Egyin Gol Valley of Mongolia". American Journal of Human Genetics. 73 (2): 247–60. doi:10.1086/377005. PMC 1180365. PMID 12858290.
- ^ Lee JY, Kuang S (2017). "A Comparative Analysis of Chinese Historical Sources and Y-DNA Studies with Regard to the Early and Medieval Turkic Peoples". Inner Asia. 19 (2): 197–239. doi:10.1163/22105018-12340089. ISSN 1464-8172. S2CID 165623743. "Analysis of the mitochondrial DNA, which is maternally inherited, shows that the Xiongnu remains from this Egyin Gol necropolis consist mainly of Asian lineages (89%). West Eurasian lineages makeup the rest (11%) (Keyser-Tracqui et al. (2003: 258). However, according to a more recent study of ancient human remains from central Mongolia, the Xiongnu population in central Mongolia possessed a higher frequency of western mitochondrial DNA haplotypes (37.5%) than the Xiongnu from the Egyin Gol necropolis (Rogers 2016: 78)."
- ^ Cai, Dawei; Zheng, Ying; Bao, Qingchuan; Hu, Xiaonong; Chen, Wenhu; Zhang, Fan; Cao, Jianen; Ning, Chao (24 November 2023). "Ancient DNA sheds light on the origin and migration patterns of the Xianbei confederation". Archaeological and Anthropological Sciences. 15 (12): 194. Bibcode:2023ArAnS..15..194C. doi:10.1007/s12520-023-01899-x. ISSN 1866-9565. S2CID 265381985.
- ^ Li et al. 2018, p. 1.
- ^ Changchun Y, Li X, Xiaolei Z, Hui Z, Hong Z (November 2006). "Genetic analysis on Tuoba Xianbei remains excavated from Qilang Mountain Cemetery in Qahar Right Wing Middle Banner of Inner Mongolia". FEBS Letters. 580 (26): 6242–6246. Bibcode:2006FEBSL.580.6242C. doi:10.1016/j.febslet.2006.10.030. PMID 17070809. S2CID 19492267.
- ^ Wang H, Ge B, Mair VH, Cai D, Xie C, Zhang Q, et al. (November 2007). "Molecular genetic analysis of remains from Lamadong cemetery, Liaoning, China". American Journal of Physical Anthropology. 134 (3): 404–411. doi:10.1002/ajpa.20685. PMID 17632796.
- ^ Zhou H (March 2014). "Genetic analyses of Xianbei populations about 1,500–1,800 years old". Human Genetics. 50 (3): 308–314. doi:10.1134/S1022795414030119. S2CID 18809679.
- ^ Osada, Naoki; Kawai, Yosuke (2021). "Exploring models of human migration to the Japanese archipelago using genome-wide genetic data". Anthropological Science. 129 (1): 45–58. doi:10.1537/ase.201215.
- ^ a b Cooke, Niall P.; Mattiangeli, Valeria; Cassidy, Lara M.; Okazaki, Kenji; Stokes, Caroline A.; Onbe, Shin; Hatakeyama, Satoshi; Machida, Kenichi; Kasai, Kenji; Tomioka, Naoto; Matsumoto, Akihiko (September 2021). "Ancient genomics reveals tripartite origins of Japanese populations". Science Advances. 7 (38): eabh2419. Bibcode:2021SciA....7.2419C. doi:10.1126/sciadv.abh2419. PMC 8448447. PMID 34533991.
- ^ Yang, Melinda A. (6 January 2022). "A genetic history of migration, diversification, and admixture in Asia". Human Population Genetics and Genomics. 2 (1): 1–32. doi:10.47248/hpgg2202010001.
- ^ 张明, 平婉菁; ZHANG Ming, PING Wanjing (15 June 2023). "古基因组揭示史前欧亚大陆现代人复杂遗传历史". 人类学学报 (in Chinese). 42 (3): 412. doi:10.16359/j.1000-3193/AAS.2023.0010. ISSN 1000-3193.
- ^ a b c d Gakuhari T, Nakagome S, Rasmussen S, Allentoft ME, Sato T, Korneliussen T, et al. (August 2020). "Ancient Jomon genome sequence analysis sheds light on migration patterns of early East Asian populations". Communications Biology. 3 (1): 437. doi:10.1038/s42003-020-01162-2. PMC 7447786. PMID 32843717.
- ^ Zhang X, He G, Li W, Wang Y, Li X, Chen Y, et al. (30 September 2021). "Genomic Insight Into the Population Admixture History of Tungusic-Speaking Manchu People in Northeast China". Frontiers in Genetics. 12: 754492. doi:10.3389/fgene.2021.754492. PMC 8515022. PMID 34659368.
 This article incorporates text available under the CC BY 4.0 license.
This article incorporates text available under the CC BY 4.0 license.
- ^ a b c Xue, Yali; Zerjal, Tatiana; Bao, Weidong; Zhu, Suling; Shu, Qunfang; Xu, Jiujin; Du, Ruofu; Fu, Songbin; Li, Pu; Hurles, Matthew E; Yang, Huanming; Tyler-Smith, Chris (1 April 2006). "Male Demography in East Asia: A North–South Contrast in Human Population Expansion Times". Genetics. 172 (4): 2431–2439. doi:10.1534/genetics.105.054270. PMC 1456369. PMID 16489223.
- ^ a b c Wang Chi-zao, Shi Mei-sen, and Li Hui, "The Origin of Daur from the Perspective of Molecular Anthropology." Journal of North Minzu University 2018, No.5, Gen.No.143.
- ^ Wei LH, Yan S, Yu G, Huang YZ, Yao DL, Li SL, et al. (March 2017). "Genetic trail for the early migrations of Aisin Gioro, the imperial house of the Qing dynasty". Journal of Human Genetics. 62 (3): 407–411. doi:10.1038/jhg.2016.142. PMID 27853133. S2CID 7685248.
- ^ Yan S, Tachibana H, Wei LH, Yu G, Wen SQ, Wang CC (June 2015). "Y chromosome of Aisin Gioro, the imperial house of the Qing dynasty". Journal of Human Genetics. 60 (6): 295–8. arXiv:1412.6274. doi:10.1038/jhg.2015.28. PMID 25833470. S2CID 7505563.
- ^ a b "Did you know DNA was used to uncover the origin of the House of Aisin Gioro?". Did You Know DNA... 14 November 2016. Retrieved 5 November 2020.
- ^ Xue Y, Zerjal T, Bao W, Zhu S, Lim SK, Shu Q, et al. (December 2005). "Recent spread of a Y-chromosomal lineage in northern China and Mongolia". American Journal of Human Genetics. 77 (6): 1112–6. doi:10.1086/498583. PMC 1285168. PMID 16380921.
- ^ "Asian Ancestry based on Studies of Y-DNA Variation: Part 3. Recent demographics and ancestry of the male East Asians – Empires and Dynasties". Genebase Tutorials. Archived from the original on 25 November 2013.
- ^ Zhang, Xianpeng; He, Guanglin; Li, Wenhui; Wang, Yunfeng; Li, Xin; Chen, Ying; Qu, Quanying; Wang, Ying; Xi, Huanjiu; Wang, Chuan-Chao; Wen, Youfeng (30 September 2021). "Genomic Insight Into the Population Admixture History of Tungusic-Speaking Manchu People in Northeast China". Frontiers in Genetics. 12: 754492. doi:10.3389/fgene.2021.754492. ISSN 1664-8021. PMC 8515022. PMID 34659368.
- ^ Hammer, Michael F.; Karafet, Tatiana M.; Park, Hwayong; Omoto, Keiichi; Harihara, Shinji; Stoneking, Mark; Horai, Satoshi (January 2006). "Dual origins of the Japanese: common ground for hunter-gatherer and farmer Y chromosomes". Journal of Human Genetics. 51 (1): 47–58. doi:10.1007/s10038-005-0322-0. PMID 16328082. S2CID 6559289.
- ^ "公益財団法人 アイヌ民族文化財団". www.ff-ainu.or.jp (in Japanese). Retrieved 8 December 2023.
- ^ Jinam, Timothy; Nishida, Nao; Hirai, Momoki; Kawamura, Shoji; Oota, Hiroki; Umetsu, Kazuo; Kimura, Ryosuke; Ohashi, Jun; Tajima, Atsushi (December 2012). "The history of human populations in the Japanese Archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations". Journal of Human Genetics. 57 (12): 787–795. doi:10.1038/jhg.2012.114. PMID 23135232.
- ^ "記者会見「日本列島3人類集団の遺伝的近縁性」". 東京大学 (in Japanese). Retrieved 8 November 2021.
- ^ a b Mitsuru Sakitani (2009). 『DNA・考古・言語の学際研究が示す新・日本列島史』 [New History of the Japanese Islands Shown by Interdisciplinary Studies on DNA, Archeology, and Language] (in Japanese). Bensei Publishing. ISBN 9784585053941.
- ^ a b Suzuki, Yuka (6 December 2012). "Ryukyuan, Ainu People Genetically Similar". Asian Scientist. Archived from the original on 16 June 2016. Retrieved 11 June 2016.
- ^ Cooke NP, Mattiangeli V, Cassidy LM, Okazaki K, Stokes CA, Onbe S, et al. (September 2021). "Ancient genomics reveals tripartite origins of Japanese populations". Science Advances. 7 (38): eabh2419. Bibcode:2021SciA....7.2419C. doi:10.1126/sciadv.abh2419. PMC 8448447. PMID 34533991.
- ^ "'Jomon woman' helps solve Japan's genetic mystery | NHK WORLD-JAPAN News". NHK WORLD. Retrieved 19 November 2020.
- ^ Jinam, Timothy; Nishida, Nao; Hirai, Momoki; Kawamura, Shoji; Oota, Hiroki; Umetsu, Kazuo; Kimura, Ryosuke; Ohashi, Jun; Tajima, Atsushi; Yamamoto, Toshimichi; Tanabe, Hideyuki; Mano, Shuhei; Suto, Yumiko; Kaname, Tadashi; Naritomi, Kenji (8 November 2012). "The history of human populations in the Japanese Archipelago inferred from genome-wide SNP data with a special reference to the Ainu and the Ryukyuan populations". Journal of Human Genetics. 57 (12): 787–795. doi:10.1038/jhg.2012.114. ISSN 1435-232X. PMID 23135232.
- ^ 弥生人DNAで迫る日本人の起源」 [The origin of Japanese people approaching with Yayoi DNA]. ja:サイエンスZERO (Television production) (in Japanese). NHK. 23 December 2018.
- ^ a b Horai, Satoshi; Murayama, Kumiko (1996). "mtDNA Polymorphism in East Asian Populations, with Special Reference to the Peopling of Japan". American Journal of Human Genetics. 59 (3). Cambridge, Massachusetts: Cell Press: 579–590. PMC 1914908. PMID 8751859.
- ^ Boer, Elisabeth de; Yang, Melinda A.; Kawagoe, Aileen; Barnes, Gina L. (2020). "Japan considered from the hypothesis of farmer/language spread". Evolutionary Human Sciences. 2: e13. doi:10.1017/ehs.2020.7. ISSN 2513-843X. PMC 10427481. PMID 37588377.
- ^ a b c Wang, Chuan-Chao (2021). "Genomic insights into the formation of human populations in East Asia". Nature. 591 (7850): 413–419. Bibcode:2021Natur.591..413W. doi:10.1038/s41586-021-03336-2. PMC 7993749. PMID 33618348.
- ^ Takeuchi F, Katsuya T, Kimura R, Nabika T, Isomura M, Ohkubo T, Tabara Y, Yamamoto K, Yokota M, Liu X, Saw WY, Mamatyusupu D, Yang W, Xu S, Teo YY, Kato N (2017). "The fine-scale genetic structure and evolution of the Japanese population". PLOS ONE. 12 (11): e0185487. Bibcode:2017PLoSO..1285487T. doi:10.1371/journal.pone.0185487. PMC 5665431. PMID 29091727.
- ^ Late Jomon male and female genome sequences from the Funadomari site in Hokkaido, Japan - Hideaki Kanzawa-Kiriyama, Department of Anthropology, National Museum of Nature and Science 2018/2019en
- ^
We also found the haplogroup M7a1 exclusively in the Korean population. This result is consistent with previous reports that haplogroup M7a is restricted to Japan and south Korea.
— Jin, et al - ^ a b Jin HJ, Tyler-Smith C, Kim W (16 January 2009). "The peopling of Korea revealed by analyses of mitochondrial DNA and Y-chromosomal markers". PLOS ONE. 4 (1): e4210. Bibcode:2009PLoSO...4.4210J. doi:10.1371/journal.pone.0004210. PMC 2615218. PMID 19148289.
- ^ Underhill PA, Shen P, Lin AA, Jin L, Passarino G, Yang WH, et al. (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics. 26 (3): 358–61. doi:10.1038/81685. PMID 11062480. S2CID 12893406.
- ^ The JPT sample is considered "as generally representative of the majority population in Japan". See Matsuda I. "Japanese in Tokyo, Japan – Population Description". Camden, NJ: Coriell Institute for Medical Research.
- ^ Poznik GD, Xue Y, Mendez FL, Willems TF, Massaia A, Wilson Sayres MA, et al. (June 2016). "Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences". Nature Genetics. 48 (6): 593–9. doi:10.1038/ng.3559. hdl:11858/00-001M-0000-002A-F024-C. PMC 4884158. PMID 27111036.
- ^ Zheng HX, Yan S, Qin ZD, Wang Y, Tan JZ, Li H, Jin L (2011). "Major population expansion of East Asians began before neolithic time: evidence of mtDNA genomes". PLOS ONE. 6 (10): e25835. Bibcode:2011PLoSO...625835Z. doi:10.1371/journal.pone.0025835. PMC 3188578. PMID 21998705.
- ^ Preucel, Robert; Mrozowski, Stephen; Nelson, Sarah (2010). Contemporary Archaeology in Theory: The New Pragmatism (2nd ed.). Wiley-Blackwell. pp. 218–221.
- ^ 水野, 文月 (15 October 2024). "弥生時代人の古代ゲノム解析から渡来人のルーツを探る". 東京大学 大学院理学系研究科・理学部 (in Japanese).
- ^ Wang & Wang 2022
- ^ Siska, Veronika; Jones, Eppie; Jeon, Sungwon; et al. (2017). "Genome-wide data from two early Neolithic East Asian individuals dating to 7700 years ago". Science Advances. 3 (2): e1601877. Bibcode:2017SciA....3E1877S. doi:10.1126/sciadv.1601877. hdl:2262/90843. PMC 5287702. PMID 28164156.
- ^ a b c Gelabert, Pere; Blazyte, Asta; Chang, Yongjoon; Fernandes, Daniel M.; Jeon, Sungwon; Hong, Jin Geun; Yoon, Jiyeon; Ko, Youngmin; Oberreiter, Victoria; Cheronet, Olivia; Özdoğan, Kadir T.; Sawyer, Susanna; Yang, Songhyok; Greytak, Ellen McRae; Choi, Hansol (8 August 2022). "Northeastern Asian and Jomon-related genetic structure in the Three Kingdoms period of Gimhae, Korea". Current Biology. 32 (15): 3232–3244.e6. Bibcode:2022CBio...32E3232G. doi:10.1016/j.cub.2022.06.004. ISSN 0960-9822. PMID 35732180.
- ^ Lee, Don-Nyeong; Jeon, Chae Lin; Kang, Jiwon; Burri, Marta; Krause, Johannes; Woo, Eun Jin; Jeong, Choongwon (December 2022). "Genomic detection of a secondary family burial in a single jar coffin in early Medieval Korea". American Journal of Biological Anthropology. 179 (4): 585–597. doi:10.1002/ajpa.24650. ISSN 2692-7691. PMC 9827920. "In both ancient and present-day Koreans, we do not detect a statistically significant contribution from the Jomon hunter-gatherer gene pool of the Japanese archipelago (Table S7A), although previous studies report occasional presence of the Jomon ancestry contribution from Neolithic to the early Medieval period (Gelabert et al., 2022; Robbeets et al., 2021). When we replace the genetic northern proxy from WLR_BA to Middle Neolithic individuals from the Miaogizou site in Inner Mongolia (“Miaozigou_MN”), we detect a small but significant amount of Jomon contribution in the Gunsan individuals and present-day Ulsan Koreans (3.1%–4.4%; Table S7B). We believe that WLR_BA provides a more suitable model for ancient and present-day Koreans given its geographical and temporal proximity to them. The remaining well-fitting source pairs provide qualitatively similar results (Table S8)."
- ^ Wang & Wang 2022: "The genetic legacy of Jomon was not restricted to Japan but was also found in Neolithic Korea."
- ^ Kim, Jungeun; Jeon, Sungwon; Choi, Jae-Pil; et al. (2020). "The Origin and Composition of Korean Ethnicity Analyzed by Ancient and Present-Day Genome Sequences". Genome Biology and Evolution. 12 (5): 553–565. doi:10.1093/gbe/evaa062. PMC 7250502. PMID 32219389.
- ^ a b c d e f Kim SH, Kim KC, Shin DJ, Jin HJ, Kwak KD, Han MS, Song JM, Kim W, Kim W (April 2011). "High frequencies of Y-chromosome haplogroup O2b-SRY465 lineages in Korea: a genetic perspective on the peopling of Korea". Investigative Genetics. 2 (1): 10. doi:10.1186/2041-2223-2-10. PMC 3087676. PMID 21463511.
 Text was copied from this source, which is available under a Creative Commons Attribution 2.0 (CC BY 2.0) license.
Text was copied from this source, which is available under a Creative Commons Attribution 2.0 (CC BY 2.0) license.
- ^ Kim SH, Han MS, Kim W, Kim W (November 2010). "Y chromosome homogeneity in the Korean population". International Journal of Legal Medicine. 124 (6): 653–7. doi:10.1007/s00414-010-0501-1. PMID 20714743. S2CID 27125545.
- ^ a b c d e So Yeun Kwon, Hwan Young Lee, Eun Young Lee, Woo Ick Yang, and Kyoung-Jin Shin, "Confirmation of Y haplogroup tree topologies with newly suggested Y-SNPs for the C2, O2b and O3a subhaplogroups." Forensic Science International: Genetics 19 (2015) 42–46. http://dx.doi.org/10.1016/j.fsigen.2015.06.003
- ^ Zhang, Ai Hua; Lee, Hye Young; Seo, Seung Bum; Lee, Hyo Jung; Jin, Hong Xuan; Cho, So Hee; Lyoo, Sung Hee; Kim, Ki Ha; Lee, Jae Won; Lee, Soong Deok (29 May 2012). "Y Haplogroup Distribution in Korean and Other Populations". Korean Journal of Legal Medicine. 36 (1): 34–44. doi:10.7580/KoreanJLegMed.2012.36.1.34.
- ^ Choi, Sun Seong; Park, Kyung Hwa; Nam, Da Eun; Kang, Tae Hoon; Chung, Ki Wha (1 December 2017). "Y-chromosome haplogrouping for Asians using Y-SNP target sequencing". Forensic Science International: Genetics Supplement Series. 6: e235 – e237. doi:10.1016/j.fsigss.2017.09.100. ISSN 1875-1768.
- ^ Kim, Soon Hee; Han, Myun Soo; Kim, Wook; Kim, Won (November 2010). "Y chromosome homogeneity in the Korean population". International Journal of Legal Medicine. 124 (6): 653–657. doi:10.1007/s00414-010-0501-1. ISSN 1437-1596. PMID 20714743. S2CID 27125545.
- ^ Jin, Han-Jun; Tyler-Smith, Chris; Kim, Wook (16 January 2009). "The Peopling of Korea Revealed by Analyses of Mitochondrial DNA and Y-Chromosomal Markers". PLOS ONE. 4 (1): e4210. Bibcode:2009PLoSO...4.4210J. doi:10.1371/journal.pone.0004210. ISSN 1932-6203. PMC 2615218. PMID 19148289.
- ^ Jeon, Sungwon; Bhak, Youngjune; Choi, Yeonsong; Jeon, Yeonsu; Kim, Seunghoon; Jang, Jaeyoung; Jang, Jinho; Blazyte, Asta; Kim, Changjae; Kim, Yeonkyung; Shim, Jungae; Kim, Nayeong; Kim, Yeo Jin; Park, Seung Gu; Kim, Jungeun (27 May 2020). "Korean Genome Project: 1094 Korean personal genomes with clinical information". Science Advances. 6 (22): eaaz7835. Bibcode:2020SciA....6.7835J. doi:10.1126/sciadv.aaz7835. ISSN 2375-2548. PMC 7385432. PMID 32766443.
- ^ Hong, Seung Beom; Kim, Ki Cheol; Kim, Wook (1 October 2014). "Mitochondrial DNA haplogroups and homogeneity in the Korean population". Genes & Genomics. 36 (5): 583–590. doi:10.1007/s13258-014-0194-9. ISSN 2092-9293. S2CID 256067815.
- ^ "Population dynamics and the rise of empires in Inner Asia: Genome-wide analysis spanning 6,000 years in the eastern Eurasian Steppe gives insights to the formation of Mongolia's empires". ScienceDaily. Retrieved 4 June 2023.
- ^ Yang, Xiaomin; Sarengaowa; He, Guanglin; Guo, Jianxin; Zhu, Kongyang; Ma, Hao; Zhao, Jing; Yang, Meiqing; Chen, Jing; Zhang, Xianpeng; Tao, Le; Liu, Yilan; Zhang, Xiu-Fang; Wang, Chuan-Chao (2021). "Genomic Insights Into the Genetic Structure and Natural Selection of Mongolians". Frontiers in Genetics. 12: 735786. doi:10.3389/fgene.2021.735786. ISSN 1664-8021. PMC 8693022. PMID 34956310.
- ^ "DNA Match Solves Ancient Mystery". china.org.cn. Retrieved 25 December 2018.
- ^ Malyarchuk BA, Derenko M, Denisova G, Woźniak M, Rogalla U, Dambueva I, Grzybowski T (June 2016). "Y chromosome haplotype diversity in Mongolic-speaking populations and gene conversion at the duplicated STR DYS385a,b in haplogroup C3-M407". Journal of Human Genetics. 61 (6): 491–496. doi:10.1038/jhg.2016.14. PMID 26911356. S2CID 13217444.
- ^ Guang‐Lin He, Meng‐Ge Wang, Xing Zou, Hui‐Yuan Yeh, Chang‐Hui Liu, Chao Liu, Gang Chen, and Chuan‐Chao Wang. "Extensive ethnolinguistic diversity at the crossroads of North China and South Siberia reflects multiple sources of genetic diversity." J Syst Evol, 2023, 61(1): 230-250. https://doi.org/10.1111/jse.12827
- ^ "Mongol Genetics – DNA of Mongolia's Khalkha Mongolians and others". khazaria.com. Retrieved 25 December 2018.
- ^ Kolman CJ, Sambuughin N, Bermingham E (April 1996). "Mitochondrial DNA analysis of Mongolian populations and implications for the origin of New World founders". Genetics. 142 (4): 1321–34. doi:10.1093/genetics/142.4.1321. PMC 1207128. PMID 8846908.
- ^ Cheng B, Tang W, He L, Dong Y, Lu J, Lei Y, et al. (October 2008). "Genetic imprint of the Mongol: signal from phylogeographic analysis of mitochondrial DNA". Journal of Human Genetics. 53 (10): 905–913. doi:10.1007/s10038-008-0325-8. PMID 18769869.
European-prevalent haplogroups (HV, U, K, I, J) are 14.3% in Xinjiang Mongolian, 10% in Mongolia, 8.4% in central Inner Mongolian samples, and only 2% in eastern Xin Barage Zuoqi County samples, showing decreasing frequencies from west to east
- ^ Feng, Qidi; Lu, Yan; Ni, X.; Yuan, K.; Yang, Ya-jun; Yang, Xiong; Liu, Chang; Lou, H.; Ning, Zhilin; Wang, Yuchen; Lu, Dongsheng; Zhang, Chao; Zhou, Ying; Shi, Meng; Tian, L. (2017). "Genetic History of Xinjiang's Uyghurs Suggests Bronze Age Multiple-Way Contacts in Eurasia". Molecular Biology and Evolution. 34 (10): 2572–2582. doi:10.1093/molbev/msx177. PMID 28595347. S2CID 28730957.
- ^ a b Wang, Yuchen; Lu Dongsheng; Chung Yeun-Jun; Xu Shuhua (2018). "Genetic structure, divergence and admixture of Han Chinese, Japanese and Korean populations". Hereditas. 155: 19. doi:10.1186/s41065-018-0057-5. PMC 5889524. PMID 29636655.
- ^ Yang, Melinda A.; Fan, Xuechun; Sun, Bo; Chen, Chungyu; Lang, Jianfeng; Ko, Ying-Chin; Tsang, Cheng-hwa; Chiu, Hunglin; Wang, Tianyi; Bao, Qingchuan; Wu, Xiaohong; Hajdinjak, Mateja; Ko, Albert Min-Shan; Ding, Manyu; Cao, Peng (17 July 2020). "Ancient DNA indicates human population shifts and admixture in northern and southern China". Science. 369 (6501): 282–288. Bibcode:2020Sci...369..282Y. doi:10.1126/science.aba0909. ISSN 0036-8075. PMID 32409524. S2CID 218649510.
- ^ Cao, Yanan; Li, Lin; Xu, Min; et al. (2020). "The ChinaMAP analytics of deep whole genome sequences in 10,588 individuals". Cell Research. 30 (9): 717–731. doi:10.1038/s41422-020-0322-9. PMC 7609296. PMID 32355288.
- ^ a b Pan, Ziqing; Xu, Shuhua (2019). "Population genomics of East Asian ethnic groups". Hereditas. 157 (49). Berlin: BioMed Central (published 2020): 49. doi:10.1186/s41065-020-00162-w. PMC 7724877. PMID 33292737.
- ^ Shi, Cheng-Min; Liu, Qi; Zhao, Shilei; Chen, Hua (21 March 2019). "Ancestry informative SNP panels for discriminating the major East Asian populations: Han Chinese, Japanese and Korean". Annals of Human Genetics. 29 (2). Cambridge: John Wiley & Sons: 348–354. doi:10.1111/ahg.12320. PMID 31025319. Archived from the original on 5 November 2021. Retrieved 14 September 2021.
- ^ a b Zhao, Yong-Bin; Zhang, Ye; Zhang, Quan-Chao; Li, Hong-Jie; Cui, Ying-Qiu; Xu, Zhi; Jin, Li; Zhou, Hui; Zhu, Hong (4 May 2015). Hofreiter, Michael (ed.). "Ancient DNA Reveals That the Genetic Structure of the Northern Han Chinese Was Shaped Prior to 3,000 Years Ago". PLOS ONE. 10 (5): e0125676. Bibcode:2015PLoSO..1025676Z. doi:10.1371/journal.pone.0125676. PMC 4418768. PMID 25938511.
- ^ Ebrey, Patricia Buckley (2023). "Rethinking Han Chinese Identity". China Review. 23 (2): 57–86. ISSN 1680-2012. JSTOR 48726991.
- ^ Feng, Qidi; Lu, Yan; Ni, X.; Yuan, K.; Yang, Ya-jun; Yang, Xiong; Liu, Chang; Lou, H.; Ning, Zhilin; Wang, Yuchen; Lu, Dongsheng; Zhang, Chao; Zhou, Ying; Shi, Meng; Tian, L. (2017). "Genetic History of Xinjiang's Uyghurs Suggests Bronze Age Multiple-Way Contacts in Eurasia". Molecular Biology and Evolution. 34 (10): 2572–2582. doi:10.1093/molbev/msx177. PMID 28595347. S2CID 28730957.
- ^ Zhao, Yong-Bin; Zhang, Ye; Zhang, Quan-Chao; Li, Hong-Jie; Cui, Ying-Qiu; Xu, Zhi; Jin, Li; Zhou, Hui; Zhu, Hong (4 May 2015). "Ancient DNA Reveals That the Genetic Structure of the Northern Han Chinese Was Shaped Prior to 3,000 Years Ago". PLOS ONE. 10 (5): e0125676. Bibcode:2015PLoSO..1025676Z. doi:10.1371/journal.pone.0125676. PMC 4418768. PMID 25938511.
- ^ Chiang, Charleston W.K.; Mangul, Serghei; Robles, Christopher; Sankararaman, Sriram (2018). "A Comprehensive Map of Genetic Variation in the World's Largest Ethnic Group —Han Chinese". Molecular Biology and Evolution. 35 (11): 2736–2750. doi:10.1093/molbev/msy170. PMC 6693441. PMID 30169787.
- ^ He, Guang-Lin; Wang, Meng-Ge; Li, Ying-Xiang; et al. (2022). "Fine-scale north-to-south genetic admixture profile in Shaanxi Han Chinese revealed by genome-wide demographic history reconstruction". Journal of Systematics and Evolution. 60 (4): 955–972. doi:10.1111/jse.12715 – via Wiley Online Library.
- ^ Zhou, Jingbin; Zhang, Xianpeng; Li, Xin; et al. (2022). "Genetic structure and demographic history of Northern Han people in Liaoning Province inferred from genome-wide array data". Frontiers in Ecology and Evolution. 10. doi:10.3389/fevo.2022.1014024.
- ^ Sun, Na; Ma, Peng-Cheng; Yan, Shi; et al. (2019). "Phylogeography of Y-chromosome haplogroup Q1a1a-M120, a paternal lineage connecting populations in Siberia and East Asia". Annals of Human Biology. 46 (3): 261–266. doi:10.1080/03014460.2019.1632930. PMID 31208219 – via Taylor & Francis Online.
- ^ Hurles, M; Sykes, B; Jobling, M; Forster, P (2005). "The Dual Origin of the Malagasy in Island Southeast Asia and East Africa: Evidence from Maternal and Paternal Lineages". The American Journal of Human Genetics. 76 (5): 894–901. doi:10.1086/430051. PMC 1199379. PMID 15793703.
- ^ a b c d e He, Guanglin; Wang, Mengge; Miao, Lei; Chen, Jing; Zhao, Jie; Sun, Qiuxia; Duan, Shuhan; Wang, Zhiyong; Xu, Xiaofei; Sun, Yuntao; Liu, Yan; Liu, Jing; Wang, Zheng; Wei, Lanhai; Liu, Chao; Ye, Jian; Wang, Le (28 March 2023). "Multiple founding paternal lineages inferred from the newly-developed 639-plex Y-SNP panel suggested the complex admixture and migration history of Chinese people". Human Genomics. 17 (1): 29. doi:10.1186/s40246-023-00476-6. PMC 10045532. PMID 36973821.
- ^ a b c Lu, Chuncheng; Zhang, Jie; Li, Yingchun; Xia, Yankai; Zhang, Feng; Wu, Bin; Wu, Wei; Ji, Guixiang; Gu, Aihua; Wang, Shoulin; Jin, Li; Wang, Xinru (2009). "The b2/b3 subdeletion shows higher risk of spermatogenic failure and higher frequency of complete AZFc deletion than the gr/gr subdeletion in a Chinese population". Human Molecular Genetics. 18 (6): 1122–30. doi:10.1093/hmg/ddn427. PMID 19088127.
- ^ a b Wen, B.; Li, H.; Lu, D.; Song, X.; Zhang, F.; He, Y.; Li, F.; Gao, Y.; et al. (September 2004). "Genetic evidence supports demic diffusion of Han culture" (PDF). Nature. 431 (7006): 302–05. Bibcode:2004Natur.431..302W. doi:10.1038/nature02878. PMID 15372031. S2CID 4301581. Archived from the original (PDF) on 24 March 2009.
- ^ Xue, Fuzhong; Wang, Yi; Xu, Shuhua; Zhang, Feng; Wen, Bo; Wu, Xuesen; Lu, Ming; Deka, Ranjan; Qian, Ji; et al. (2008). "A spatial analysis of genetic structure of human populations in China reveals distinct difference between maternal and paternal lineages". European Journal of Human Genetics. 16 (6): 705–17. doi:10.1038/sj.ejhg.5201998. PMID 18212820.
- ^ Wen, Bo; Li, Hui; Lu, Daru; Song, Xiufeng; Zhang, Feng; He, Yungang; Li, Feng; Gao, Yang; Mao, Xianyun; et al. (2004). "Genetic evidence supports demic diffusion of Han culture". Nature. 431 (7006): 302–05. Bibcode:2004Natur.431..302W. doi:10.1038/nature02878. PMID 15372031. S2CID 4301581.
- ^ Chen, Jieming; Zheng, Houfeng; Bei, Jin-Xin; Sun, Liangdan; Jia, Wei-hua; Li, Tao; Zhang, Furen; Seielstad, Mark; Zeng, Yi-Xin; et al. (2009). "Genetic Structure of the Han Chinese Population Revealed by Genome-wide SNP Variation". The American Journal of Human Genetics. 85 (6): 775–85. doi:10.1016/j.ajhg.2009.10.016. PMC 2790583. PMID 19944401.
- ^ McFadzean A.J.S., Todd D. (1971). "Cooley's anaemia among the tanka of South China". Transactions of the Royal Society of Tropical Medicine and Hygiene. 65 (1): 59–62. doi:10.1016/0035-9203(71)90185-4. PMID 5092429.
- ^ Gan, Rui-Jing; Pan, Shang-Ling; Mustavich, Laura F.; Qin, Zhen-Dong; Cai, Xiao-Yun; Qian, Ji; Liu, Cheng-Wu; Peng, Jun-Hua; Li, Shi-Lin; Xu, Jie-Shun; Jin, Li; Li, Hui (2008). "Pinghua population as an exception of Han Chinese's coherent genetic structure". Journal of Human Genetics. 53 (4): 303–13. doi:10.1007/s10038-008-0250-x. PMID 18270655.
- ^ Yang, Xiaomin; Wang, Xiao-Xun; He, Guanglin; et al. (2020). "Genomic insight into the population history of Central Han Chinese". Annals of Human Biology – via ResearchGate.
- ^ Wang Y, Lu D, Chung YJ, Xu S (6 April 2018). "Genetic structure, divergence and admixture of Han Chinese, Japanese and Korean populations". Hereditas. 155 (1): 19. doi:10.1186/s41065-018-0057-5. PMC 5889524. PMID 29636655. *Xu S (10 April 2018). "Common ancestor of Han Chinese, Japanese and Koreans dated to 3000 – 3600 years ago". Biomed Central.
- ^ Cao Y, Li L, Xu M, Feng Z, Sun X, Lu J, et al. (September 2020). "The ChinaMAP analytics of deep whole genome sequences in 10,588 individuals". Cell Research. 30 (9): 717–731. doi:10.1038/s41422-020-0322-9. PMC 7609296. PMID 32355288.
- ^ a b Zhao YB, Zhang Y, Zhang QC, Li HJ, Cui YQ, Xu Z, Jin L, Zhou H, Zhu H (2015). "Ancient DNA reveals that the genetic structure of the northern Han Chinese was shaped prior to 3,000 years ago". PLOS ONE. 10 (5): e0125676. Bibcode:2015PLoSO..1025676Z. doi:10.1371/journal.pone.0125676. PMC 4418768. PMID 25938511.
- ^ Li J, Zeng W, Zhang Y, Ko AM, Li C, Zhu H, Fu Q, Zhou H (December 2017). "Ancient DNA reveals genetic connections between early Di-Qiang and Han Chinese". BMC Evolutionary Biology. 17 (1): 239. Bibcode:2017BMCEE..17..239L. doi:10.1186/s12862-017-1082-0. PMC 5716020. PMID 29202706.
- ^ Yang MA, Fan X, Sun B, Chen C, Lang J, Ko YC, et al. (July 2020). "Ancient DNA indicates human population shifts and admixture in northern and southern China". Science. 369 (6501): 282–288. Bibcode:2020Sci...369..282Y. doi:10.1126/science.aba0909. PMID 32409524. S2CID 218649510.
- ^ a b Wen B, Li H, Lu D, Song X, Zhang F, He Y, et al. (September 2004). "Genetic evidence supports demic diffusion of Han culture". Nature. 431 (7006): 302–305. Bibcode:2004Natur.431..302W. doi:10.1038/nature02878. PMID 15372031. S2CID 4301581.
- ^ Du R, Xiao C, Cavalli-Sforza LL (December 1997). "Genetic distances between Chinese populations calculated on gene frequencies of 38 loci". Science in China Series C: Life Sciences. 40 (6): 613–21. doi:10.1007/BF02882691. PMID 18726285. S2CID 1924085.
- ^ Qin, Pengfei; Zhou, Ying; Lou, Haiyi; Lu, Dongsheng; Yang, Xiong; Wang, Yuchen; Jin, Li; Chung, Yeun-Jun; Xu, Shuhua (2 April 2015). "Quantitating and Dating Recent Gene Flow between European and East Asian Populations". Scientific Reports. 5 (1): 9500. Bibcode:2015NatSR...5.9500Q. doi:10.1038/srep09500. ISSN 2045-2322. PMC 4382708. PMID 25833680.
- ^ Chen J, Zheng H, Bei JX, Sun L, Jia WH, Li T, et al. (December 2009). "Genetic structure of the Han Chinese population revealed by genome-wide SNP variation". American Journal of Human Genetics. 85 (6): 775–785. doi:10.1016/j.ajhg.2009.10.016. PMC 2790583. PMID 19944401.
- ^ Gan RJ, Pan SL, Mustavich LF, Qin ZD, Cai XY, Qian J, Liu CW, Peng JH, Li SL, Xu JS, Jin L, Li H (2008). "Pinghua population as an exception of Han Chinese's coherent genetic structure". Journal of Human Genetics. 53 (4): 303–13. doi:10.1007/s10038-008-0250-x. PMID 18270655.
- ^ a b He, Guanglin; Wang, Mengge; Miao, Lei; Chen, Jing; Zhao, Jie; Sun, Qiuxia; Duan, Shuhan; Wang, Zhiyong; Xu, Xiaofei; Sun, Yuntao; Liu, Yan; Liu, Jing; Wang, Zheng; Wei, Lanhai; Liu, Chao (28 March 2023). "Multiple founding paternal lineages inferred from the newly-developed 639-plex Y-SNP panel suggested the complex admixture and migration history of Chinese people". Human Genomics. 17 (1): 29. doi:10.1186/s40246-023-00476-6. ISSN 1479-7364. PMC 10045532. PMID 36973821.
- ^ Yao YG, Kong QP, Bandelt HJ, Kivisild T, Zhang YP (March 2002). "Phylogeographic differentiation of mitochondrial DNA in Han Chinese". American Journal of Human Genetics. 70 (3): 635–51. doi:10.1086/338999. PMC 384943. PMID 11836649.
- ^ Kivisild T, Tolk HV, Parik J, Wang Y, Papiha SS, Bandelt HJ, Villems R (October 2002). "The emerging limbs and twigs of the East Asian mtDNA tree". Molecular Biology and Evolution. 19 (10): 1737–51. doi:10.1093/oxfordjournals.molbev.a003996. PMID 12270900.
- ^ Yao YG, Kong QP, Man XY, Bandelt HJ, Zhang YP (February 2003). "Reconstructing the evolutionary history of China: a caveat about inferences drawn from ancient DNA". Molecular Biology and Evolution. 20 (2): 214–9. doi:10.1093/molbev/msg026. PMID 12598688.
- ^ Kong QP, Sun C, Wang HW, Zhao M, Wang WZ, Zhong L, Hao XD, Pan H, Wang SY, Cheng YT, Zhu CL, Wu SF, Liu LN, Jin JQ, Yao YG, Zhang YP (January 2011). "Large-scale mtDNA screening reveals a surprising matrilineal complexity in east Asia and its implications to the peopling of the region". Molecular Biology and Evolution. 28 (1): 513–22. doi:10.1093/molbev/msq219. PMID 20713468.
- ^ Liu, Chi-Chun; Witonsky, David; Gosling, Anna; Lee, Ju Hyeon; Ringbauer, Harald; Hagan, Richard; Patel, Nisha; Stahl, Raphaela; Novembre, John; Aldenderfer, Mark; Warinner, Christina; Di Rienzo, Anna; Jeong, Choongwon (8 March 2022). "Ancient genomes from the Himalayas illuminate the genetic history of Tibetans and their Tibeto-Burman speaking neighbors". Nature Communications. 13 (1): 1203. Bibcode:2022NatCo..13.1203L. doi:10.1038/s41467-022-28827-2. ISSN 2041-1723. PMC 8904508. PMID 35260549. S2CID 247317520.
- ^ Wang, Hongru; Yang, Melinda A.; Wangdue, Shargan; Lu, Hongliang; Chen, Honghai; Li, Linhui; Dong, Guanghui; Tsring, Tinley; Yuan, Haibing; He, Wei; Ding, Manyu; Wu, Xiaohong; Li, Shuai; Tashi, Norbu; Yang, Tsho (15 March 2023). "Human genetic history on the Tibetan Plateau in the past 5100 years". Science Advances. 9 (11): eadd5582. Bibcode:2023SciA....9D5582W. doi:10.1126/sciadv.add5582. ISSN 2375-2548. PMC 10022901. PMID 36930720.
- ^ Bhandari, Sushil; Zhang, Xiaoming (5 November 2015). "Genetic evidence of a recent Tibetan ancestry to Sherpas in the Himalayan region". Scientific Reports. 5: 16249. Bibcode:2015NatSR...516249B. doi:10.1038/srep16249. ISSN 2045-2322. PMC 4633682. PMID 26538459. "Comparing Sherpas, Tibetans, and Han Chinese showed that the D-M174 is the predominant haplogroup in Sherpas (43.38%) and prevalent in Tibetans (52.84%)5, but rare among both Han Chinese (1.4–6.51%)6,7 and other Asian populations (0.02–0.07%)8, aside from Japanese (34.7%) who possesses a distinct D-M174 lineage highly diverged from those in Tibetans and other Asian populations9,10."
- ^ Zhang, Ganyu; Cui, Can; Wangdue, Shargan (16 March 2023). "Maternal genetic history of ancient Tibetans over the past 4000 years". Journal of Genetics and Genomics. 50 (10): 765–775. doi:10.1016/j.jgg.2023.03.007. PMID 36933795. S2CID 257588399.
- ^ Blench R, Spriggs M (2 September 2003). Archaeology and Language II: Archaeological Data and Linguistic Hypotheses. Routledge. ISBN 978-1-134-82869-2.
- ^ Yunusbayev, Bayazit; Metspalu, Mait; Metspalu, Ene; Valeev, Albert; Litvinov, Sergei; Valiev, Ruslan; Akhmetova, Vita; Balanovska, Elena; Balanovsky, Oleg; Turdikulova, Shahlo; Dalimova, Dilbar; Nymadawa, Pagbajabyn; Bahmanimehr, Ardeshir; Sahakyan, Hovhannes; Tambets, Kristiina (21 April 2015). "The Genetic Legacy of the Expansion of Turkic-Speaking Nomads across Eurasia". PLOS Genetics. 11 (4): e1005068. doi:10.1371/journal.pgen.1005068. ISSN 1553-7404. PMC 4405460. PMID 25898006.
- ^ Nelson S, Zhushchikhovskaya I, Li T, Hudson M, Robbeets M (2020). "Tracing population movements in ancient East Asia through the linguistics and archaeology of textile production". Evolutionary Human Sciences. 2: e5. doi:10.1017/ehs.2020.4. ISSN 2513-843X. PMC 10427276. PMID 37588355. S2CID 213436897.
- ^ Li T, Ning C, Zhushchikhovskaya IS, Hudson MJ, Robbeets M (1 June 2020). "Millet agriculture dispersed from Northeast China to the Russian Far East: Integrating archaeology, genetics, and linguistics". Archaeological Research in Asia. 22: 100177. doi:10.1016/j.ara.2020.100177. hdl:21.11116/0000-0005-D82B-8. ISSN 2352-2267. S2CID 213952845.
- ^ Uchiyama J, Gillam JC, Savelyev A, Ning C (2020). "Populations dynamics in Northern Eurasian forests: a long-term perspective from Northeast Asia". Evolutionary Human Sciences. 2: e16. doi:10.1017/ehs.2020.11. ISSN 2513-843X. PMC 10427466. PMID 37588381. S2CID 219470000.
- ^ Golden, Peter B. (October 2018). "The Ethnogonic Tales of the Türks". The Medieval History Journal. 21 (2): 291–327. doi:10.1177/0971945818775373. ISSN 0971-9458. S2CID 166026934.
- ^ Dai, Shan-Shan; Sulaiman, Xierzhatijiang; Isakova, Jainagul; Xu, Wei-Fang; Abdulloevich, Najmudinov Tojiddin; Afanasevna, Manilova Elena; Ibrohimovich, Khudoidodov Behruz; Chen, Xi; Yang, Wei-Kang; Wang, Ming-Shan; Shen, Quan-Kuan; Yang, Xing-Yan; Yao, Yong-Gang; Aldashev, Almaz A; Saidov, Abdusattor (25 August 2022). "The Genetic Echo of the Tarim Mummies in Modern Central Asians". Molecular Biology and Evolution. 39 (9). doi:10.1093/molbev/msac179. ISSN 0737-4038. PMC 9469894. PMID 36006373.
- ^ Guarino-Vignon, Perle; Marchi, Nina; Bendezu-Sarmiento, Julio; Heyer, Evelyne; Bon, Céline (14 January 2022). "Genetic continuity of Indo-Iranian speakers since the Iron Age in southern Central Asia". Scientific Reports. 12 (1): 733. Bibcode:2022NatSR..12..733G. doi:10.1038/s41598-021-04144-4. ISSN 2045-2322. PMC 8760286. PMID 35031610.
- ^ Zerjal T, Wells RS, Yuldasheva N, Ruzibakiev R, Tyler-Smith C (September 2002). "A genetic landscape reshaped by recent events: Y-chromosomal insights into central Asia". American Journal of Human Genetics. 71 (3): 466–82. doi:10.1086/342096. PMC 419996. PMID 12145751.
- ^ Kumar D (11 May 2012). Genomics and Health in the Developing World. Oxford, England: Oxford University Press. pp. 1265–1267. ISBN 978-0-19-970547-4.
- ^ He, Guanglin; Wang, Mengge; Luo, Lintao; Sun, Qiuxia; Yuan, Haibing; Lv, Hongliang; Feng, Yuhang; Liu, Xiaojun; Cheng, Jing; Bu, Fengxiao; Zhabagin, Maxat; Yuan, Huijun; Liu, Chao; Xu, Shuhua (4 November 2024). "Population genomics of Central Asian peoples unveil ancient Trans-Eurasian genetic admixture and cultural exchanges". hLife. 2 (11): 554–562. doi:10.1016/j.hlife.2024.06.006. ISSN 2949-9283.
- ^ Damgaard PB, Marchi N, Rasmussen S, Peyrot M, Renaud G, Korneliussen T, et al. (May 2018). "137 ancient human genomes from across the Eurasian steppes". Nature. 557 (7705): 369–374. Bibcode:2018Natur.557..369D. doi:10.1038/s41586-018-0094-2. hdl:1887/3202709. PMID 29743675. S2CID 13670282.
- ^ Yunusbayev B, Metspalu M, Metspalu E, Valeev A, Litvinov S, Valiev R, et al. (April 2015). "The genetic legacy of the expansion of Turkic-speaking nomads across Eurasia". PLOS Genetics. 11 (4): e1005068. doi:10.1371/journal.pgen.1005068. PMC 4405460. PMID 25898006.
- ^ Moreno-Mayar JV, Potter BA, Vinner L, Steinrücken M, Rasmussen S, Terhorst J, et al. (January 2018). "Terminal Pleistocene Alaskan genome reveals first founding population of Native Americans" (PDF). Nature. 553 (7687): 203–207. Bibcode:2018Natur.553..203M. doi:10.1038/nature25173. PMID 29323294. S2CID 4454580.
- ^ Davis LG, Madsen DB, Becerra-Valdivia L, Higham T, Sisson DA, Skinner SM, et al. (August 2019). "Late Upper Paleolithic occupation at Cooper's Ferry, Idaho, USA, ~16,000 years ago". Science. 365 (6456): 891–897. Bibcode:2019Sci...365..891D. doi:10.1126/science.aax9830. PMID 31467216.
- ^ Posth C, Nakatsuka N, Lazaridis I, Skoglund P, Mallick S, Lamnidis TC, et al. (November 2018). "Reconstructing the Deep Population History of Central and South America". Cell. 175 (5): 1185–1197.e22. doi:10.1016/j.cell.2018.10.027. PMC 6327247. PMID 30415837.
- ^ Willerslev E, Meltzer DJ (June 2021). "Peopling of the Americas as inferred from ancient genomics". Nature. 594 (7863): 356–364. Bibcode:2021Natur.594..356W. doi:10.1038/s41586-021-03499-y. PMID 34135521. S2CID 235460793.
- ^ Sarkar AA (18 June 2021). "Ancient Human Genomes Reveal Peopling of the Americas". GEN – Genetic Engineering and Biotechnology News. Retrieved 15 September 2021.
The team discovered that the Spirit Cave remains came from a Native American while dismissing a longstanding theory that a group called Paleoamericans existed in North America before Native Americans.
- ^ Yelmen B, Mondal M, Marnetto D, Pathak AK, Montinaro F, Gallego Romero I, et al. (August 2019). "Ancestry-Specific Analyses Reveal Differential Demographic Histories and Opposite Selective Pressures in Modern South Asian Populations". Molecular Biology and Evolution. 36 (8): 1628–1642. doi:10.1093/molbev/msz037. PMC 6657728. PMID 30952160.
- ^ a b Chaubey G (January 2015). "East Asian ancestry in India" (PDF). Indian Journal of Physical Anthropology and Human Genetics. 34 (2): 193–199.
Here the analysis of genome wide data on Indian and East/Southeast Asian demonstrated their restricted distinctive ancestry in India mainly running along the foothills of Himalaya and northeastern part.
- ^ a b Chaubey G, Metspalu M, Choi Y, Mägi R, Romero IG, Soares P, et al. (February 2011). "Population genetic structure in Indian Austroasiatic speakers: the role of landscape barriers and sex-specific admixture". Molecular Biology and Evolution. 28 (2): 1013–1024. doi:10.1093/molbev/msq288. PMC 3355372. PMID 20978040.
- ^ Ness I (2014). The Global Prehistory of Human Migration. Chichester, West Sussex: John Wiley & Sons. p. 265. ISBN 978-1-118-97058-4.
- ^ a b Zhang X, Liao S, Qi X, Liu J, Kampuansai J, Zhang H, Yang Z, Serey B, Sovannary T, Bunnath L, Seang Aun H, Samnom H, Kangwanpong D, Shi H, Su B (October 2015). "Y-chromosome diversity suggests southern origin and Paleolithic backwave migration of Austro-Asiatic speakers from eastern Asia to the Indian subcontinent". Scientific Reports. 5: 15486. Bibcode:2015NatSR...515486Z. doi:10.1038/srep15486. PMC 4611482. PMID 26482917.
- ^ Chaubey G, Metspalu M, Choi Y, Mägi R, Romero IG, Soares P, et al. (February 2011). "Population genetic structure in Indian Austroasiatic speakers: the role of landscape barriers and sex-specific admixture". Molecular Biology and Evolution. 28 (2): 1013–1024. doi:10.1093/molbev/msq288. PMC 3355372. PMID 20978040.
- ^ a b Van Driem G (2007). "Austroasiatic phylogeny and the Austroasiatic homeland in light of recent population genetic studies" (PDF). Mon-Khmer Studies. 37: 1–14.[dead link]
- ^ Riccio ME, Nunes JM, Rahal M, Kervaire B, Tiercy JM, Sanchez-Mazas A (June 2011). "The Austroasiatic Munda population from India and Its enigmatic origin: a HLA diversity study". Human Biology. 83 (3): 405–35. doi:10.3378/027.083.0306. PMID 21740156. S2CID 39428816.
- ^ Arunkumar G, Wei LH, Kavitha VJ, Syama A, Arun VS, Sathua S, et al. (November 2015). "A late Neolithic expansion of Y chromosomal haplogroup O2a1-M95 from east to west: Late Neolithic expansion of O2a1-M95". Journal of Systematics and Evolution. 53 (6): 546–560. doi:10.1111/jse.12147. S2CID 83103649.
- ^ "ASI-AAA"
- ^ Lévi S, Przyluski J, Bloch J (1993). Pre-Aryan and Pre-Dravidian in India. Asian Educational Services. ISBN 978-81-206-0772-9.
It has been further proved that not only linguistic but also certain cultural and political facts of ancient India, can be explained by Austroasiatic elements.
- ^ "How rice farming may have spread across the ancient world". science.org. Retrieved 26 October 2021.
- ^ Changmai P, Pinhasi R, Pietrusewsky M, Stark MT, Ikehara-Quebral RM, Reich D, Flegontov P (December 2022). "Ancient DNA from Protohistoric Period Cambodia indicates that South Asians admixed with local populations as early as 1st-3rd centuries CE". Scientific Reports. 12 (1): 22507. Bibcode:2022NatSR..1222507C. doi:10.1038/s41598-022-26799-3. PMC 9800559. PMID 36581666.
- ^ Liu D, Duong NT, Ton ND, Van Phong N, Pakendorf B, Van Hai N, Stoneking M (September 2020). "Extensive Ethnolinguistic Diversity in Vietnam Reflects Multiple Sources of Genetic Diversity". Molecular Biology and Evolution. 37 (9): 2503–2519. doi:10.1093/molbev/msaa099. PMC 7475039. PMID 32344428.
- ^ Carlhoff S, Duli A, Nägele K, Nur M, Skov L, Sumantri I, et al. (August 2021). "Genome of a middle Holocene hunter-gatherer from Wallacea". Nature. 596 (7873): 543–547. Bibcode:2021Natur.596..543C. doi:10.1038/s41586-021-03823-6. hdl:10072/407535. PMC 8387238. PMID 34433944.
The qpGraph analysis confirmed this branching pattern, with the Leang Panninge individual branching off from the Near Oceanian clade after the Denisovan gene flow, although with the most supported topology indicating around 50% of a basal East Asian component contributing to the Leang Panninge genome (Fig. 3c, Supplementary Figs. 7–11).
- ^ Changmai P, Jaisamut K, Kampuansai J, Kutanan W, Altınışık NE, Flegontova O, et al. (February 2022). "Indian genetic heritage in Southeast Asian populations". PLOS Genetics. 18 (2): e1010036. doi:10.1371/journal.pgen.1010036. PMC 8853555. PMID 35176016.
At 12 hypothetical ancestral populations, Mon, Khmer from Thailand, Kuy, Nyahkur, Burmese, Thai, Cambodian Khmer, Cham, Ede, Giarai, and Malay... demonstrated a "light pink" ancestry component (accounting for more than 5% of their ancestry, on average) that is enriched in South Asian populations such as Irula and Mala from Southern India (Fig 3)
- ^ Yang, Melinda A. (6 January 2022). "A genetic history of migration, diversification, and admixture in Asia". Human Population Genetics and Genomics. 2 (1): 1–32. doi:10.47248/hpgg2202010001. ISSN 2770-5005.
Works cited
[edit]- Lazaridis I, Belfer-Cohen A, Mallick S, Patterson N, Cheronet O, Rohland N, et al. (21 September 2018). "Paleolithic DNA from the Caucasus reveals core of West Eurasian ancestry". bioRxiv 10.1101/423079.
- Lee J, Miller BK, Bayarsaikhan J, Johannesson E, Ventresca Miller A, Warinner C, Jeong C (April 2023). "Genetic population structure of the Xiongnu Empire at imperial and local scales". Science Advances. 9 (15): eadf3904. Bibcode:2023SciA....9F3904L. doi:10.1126/sciadv.adf3904. PMC 10104459. PMID 37058560.
- Li J, Zhang Y, Zhao Y, Chen Y, Ochir A, Zhu H, Zhou H (August 2018). "The genome of an ancient Rouran individual reveals an important paternal lineage in the Donghu population". American Journal of Physical Anthropology. 166 (4). American Association of Physical Anthropologists: 895–905. doi:10.1002/ajpa.23491. PMID 29681138.
Further reading
[edit]- He G, Wang Z, Guo J, Wang M, Zou X, Tang R, et al. (August 2020). "Inferring the population history of Tai-Kadai-speaking people and southernmost Han Chinese on Hainan Island by genome-wide array genotyping". European Journal of Human Genetics. 28 (8): 1111–1123. doi:10.1038/s41431-020-0599-7. PMC 7381617. PMID 32123326. S2CID 211729663.
 KSF
KSF

