Protein
 From Wikipedia - Reading time: 41 min
From Wikipedia - Reading time: 41 min
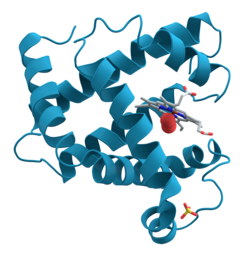
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity.
A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residues in a protein is defined by the sequence of a gene, which is encoded in the genetic code. In general, the genetic code specifies 20 standard amino acids; but in certain organisms the genetic code can include selenocysteine and—in certain archaea—pyrrolysine. Shortly after or even during synthesis, the residues in a protein are often chemically modified by post-translational modification, which alters the physical and chemical properties, folding, stability, activity, and ultimately, the function of the proteins. Some proteins have non-peptide groups attached, which can be called prosthetic groups or cofactors. Proteins can work together to achieve a particular function, and they often associate to form stable protein complexes.
Once formed, proteins only exist for a certain period and are then degraded and recycled by the cell's machinery through the process of protein turnover. A protein's lifespan is measured in terms of its half-life and covers a wide range. They can exist for minutes or years with an average lifespan of 1–2 days in mammalian cells. Abnormal or misfolded proteins are degraded more rapidly either due to being targeted for destruction or due to being unstable.
Like other biological macromolecules such as polysaccharides and nucleic acids, proteins are essential parts of organisms and participate in virtually every process within cells. Many proteins are enzymes that catalyse biochemical reactions and are vital to metabolism. Some proteins have structural or mechanical functions, such as actin and myosin in muscle, and the cytoskeleton's scaffolding proteins that maintain cell shape. Other proteins are important in cell signaling, immune responses, cell adhesion, and the cell cycle. In animals, proteins are needed in the diet to provide the essential amino acids that cannot be synthesized. Digestion breaks the proteins down for metabolic use.
History and etymology
[edit]Discovery and early studies
[edit]Proteins have been studied and recognized since the 1700s by Antoine Fourcroy and others,[1][2] who often collectively called them "albumins", or "albuminous materials" (Eiweisskörper, in German).[2] Gluten, for example, was first separated from wheat in published research around 1747, and later determined to exist in many plants.[1] In 1789, Antoine Fourcroy recognized three distinct varieties of animal proteins: albumin, fibrin, and gelatin.[3] Vegetable (plant) proteins studied in the late 1700s and early 1800s included gluten, plant albumin, gliadin, and legumin.[1]
Proteins were first described by the Dutch chemist Gerardus Johannes Mulder and named by the Swedish chemist Jöns Jacob Berzelius in 1838.[4][5] Mulder carried out elemental analysis of common proteins and found that nearly all proteins had the same empirical formula, C400H620N100O120P1S1.[6] He came to the erroneous conclusion that they might be composed of a single type of (very large) molecule. The term "protein" to describe these molecules was proposed by Mulder's associate Berzelius; protein is derived from the Greek word πρώτειος (proteios), meaning "primary",[7] "in the lead", or "standing in front",[2] + -in. Mulder went on to identify the products of protein degradation such as the amino acid leucine for which he found a (nearly correct) molecular weight of 131 Da.[6]
Early nutritional scientists such as the German Carl von Voit believed that protein was the most important nutrient for maintaining the structure of the body, because it was generally believed that "flesh makes flesh".[8] Around 1862, Karl Heinrich Ritthausen isolated the amino acid glutamic acid.[9] Thomas Burr Osborne compiled a detailed review of the vegetable proteins at the Connecticut Agricultural Experiment Station. Osborne, alongside Lafayette Mendel, established several nutritionally essential amino acids in feeding experiments with laboratory rats.[10] Diets lacking an essential amino acid stunts the rats' growth, consistent with Liebig's law of the minimum.[11] The final essential amino acid to be discovered, threonine, was identified by William Cumming Rose.[12]
The difficulty in purifying proteins impeded work by early protein biochemists. Proteins could be obtained in large quantities from blood, egg whites, and keratin, but individual proteins were unavailable. In the 1950s, the Armour Hot Dog Company purified 1 kg of bovine pancreatic ribonuclease A and made it freely available to scientists. This gesture helped ribonuclease A become a major target for biochemical study for the following decades.[6]
Polypeptides
[edit]
The understanding of proteins as polypeptides, or chains of amino acids, came through the work of Franz Hofmeister and Hermann Emil Fischer in 1902.[13][14] The central role of proteins as enzymes in living organisms that catalyzed reactions was not fully appreciated until 1926, when James B. Sumner showed that the enzyme urease was in fact a protein.[15]
Linus Pauling is credited with the successful prediction of regular protein secondary structures based on hydrogen bonding, an idea first put forth by William Astbury in 1933.[16] Later work by Walter Kauzmann on denaturation,[17][18] based partly on previous studies by Kaj Linderstrøm-Lang,[19] contributed an understanding of protein folding and structure mediated by hydrophobic interactions.[20]
The first protein to have its amino acid chain sequenced was insulin, by Frederick Sanger, in 1949. Sanger correctly determined the amino acid sequence of insulin, thus conclusively demonstrating that proteins consisted of linear polymers of amino acids rather than branched chains, colloids, or cyclols.[21] He won the Nobel Prize for this achievement in 1958.[22] Christian Anfinsen's studies of the oxidative folding process of ribonuclease A, for which he won the nobel prize in 1972, solidified the thermodynamic hypothesis of protein folding, according to which the folded form of a protein represents its free energy minimum.[23][24]
Structure
[edit]
With the development of X-ray crystallography, it became possible to determine protein structures as well as their sequences.[25] The first protein structures to be solved were hemoglobin by Max Perutz and myoglobin by John Kendrew, in 1958.[26][27] The use of computers and increasing computing power has supported the sequencing of complex proteins. In 1999, Roger Kornberg sequenced the highly complex structure of RNA polymerase using high intensity X-rays from synchrotrons.[25]
Since then, cryo-electron microscopy (cryo-EM) of large macromolecular assemblies[28] has been developed. Cryo-EM uses protein samples that are frozen rather than crystals, and beams of electrons rather than X-rays. It causes less damage to the sample, allowing scientists to obtain more information and analyze larger structures.[25] Computational protein structure prediction of small protein structural domains[29] has helped researchers to approach atomic-level resolution of protein structures. As of April 2024[update], the Protein Data Bank contains 181,018 X-ray, 19,809 EM and 12,697 NMR protein structures.[30]
Classification
[edit]Proteins are primarily classified by sequence and structure, although other classifications are commonly used. Especially for enzymes the EC number system provides a functional classification scheme.[31] Similarly, gene ontology classifies both genes and proteins by their biological and biochemical function, and by their intracellular location.[32]
Sequence similarity is used to classify proteins both in terms of evolutionary and functional similarity. This may use either whole proteins or protein domains, especially in multi-domain proteins. Protein domains allow protein classification by a combination of sequence, structure and function, and they can be combined in many ways. In an early study of 170,000 proteins, about two-thirds were assigned at least one domain, with larger proteins containing more domains (e.g. proteins larger than 600 amino acids having an average of more than 5 domains).[33]
Biochemistry
[edit]

Most proteins consist of linear polymers built from series of up to 20 L-α-amino acids. All proteinogenic amino acids have a common structure where an α-carbon is bonded to an amino group, a carboxyl group, and a variable side chain. Only proline differs from this basic structure as its side chain is cyclical, bonding to the amino group, limiting protein chain flexibility.[34] The side chains of the standard amino acids have a variety of chemical structures and properties, and it is the combined effect of all amino acids that determines its three-dimensional structure and chemical reactivity.[35]
The amino acids in a polypeptide chain are linked by peptide bonds between amino and carboxyl group. An individual amino acid in a chain is called a residue, and the linked series of carbon, nitrogen, and oxygen atoms are known as the main chain or protein backbone.[36]: 19 The peptide bond has two resonance forms that confer some double-bond character to the backbone. The alpha carbons are roughly coplanar with the nitrogen and the carbonyl (C=O) group. The other two dihedral angles in the peptide bond determine the local shape assumed by the protein backbone. One consequence of the N-C(O) double bond character is that proteins are somewhat rigid.[36]: 31 A polypeptide chain ends with a free amino group, known as the N-terminus or amino terminus, and a free carboxyl group, known as the C-terminus or carboxy terminus.[37] By convention, peptide sequences are written N-terminus to C-terminus, correlating with the order in which proteins are synthesized by ribosomes.[37][38]
The words protein, polypeptide, and peptide are a little ambiguous and can overlap in meaning. Protein is generally used to refer to the complete biological molecule in a stable conformation, whereas peptide is generally reserved for a short amino acid oligomers often lacking a stable 3D structure. But the boundary between the two is not well defined and usually lies near 20–30 residues.[39]
Proteins can interact with many types of molecules and ions, including with other proteins, with lipids, with carbohydrates, and with DNA.[40][41][42]
Abundance in cells
[edit]A typical bacterial cell, e.g. E. coli and Staphylococcus aureus, is estimated to contain about 2 million proteins. Smaller bacteria, such as Mycoplasma or spirochetes contain fewer molecules, on the order of 50,000 to 1 million. By contrast, eukaryotic cells are larger and thus contain much more protein. For instance, yeast cells have been estimated to contain about 50 million proteins and human cells on the order of 1 to 3 billion.[43] The concentration of individual protein copies ranges from a few molecules per cell up to 20 million.[44] Not all genes coding proteins are expressed in most cells and their number depends on, for example, cell type and external stimuli. For instance, of the 20,000 or so proteins encoded by the human genome, only 6,000 are detected in lymphoblastoid cells.[45] The most abundant protein in nature is thought to be RuBisCO, an enzyme that catalyzes the incorporation of carbon dioxide into organic matter in photosynthesis. Plants can consist of as much as 1% by weight of this enzyme.[46]
Synthesis
[edit]Biosynthesis
[edit]
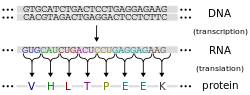
Proteins are assembled from amino acids using information encoded in genes. Each protein has its own unique amino acid sequence that is specified by the nucleotide sequence of the gene encoding this protein. The genetic code is a set of three-nucleotide sets called codons and each three-nucleotide combination designates an amino acid, for example AUG (adenine–uracil–guanine) is the code for methionine. Because DNA contains four nucleotides, the total number of possible codons is 64; hence, there is some redundancy in the genetic code, with some amino acids specified by more than one codon.[42]: 1002–42 Genes encoded in DNA are first transcribed into pre-messenger RNA (mRNA) by proteins such as RNA polymerase. Most organisms then process the pre-mRNA (a primary transcript) using various forms of post-transcriptional modification to form the mature mRNA, which is then used as a template for protein synthesis by the ribosome. In prokaryotes the mRNA may either be used as soon as it is produced, or be bound by a ribosome after having moved away from the nucleoid. In contrast, eukaryotes make mRNA in the cell nucleus and then translocate it across the nuclear membrane into the cytoplasm, where protein synthesis then takes place. The rate of protein synthesis is higher in prokaryotes than eukaryotes and can reach up to 20 amino acids per second.[47]
The process of synthesizing a protein from an mRNA template is known as translation. The mRNA is loaded onto the ribosome and is read three nucleotides at a time by matching each codon to its base pairing anticodon located on a transfer RNA molecule, which carries the amino acid corresponding to the codon it recognizes. The enzyme aminoacyl tRNA synthetase "charges" the tRNA molecules with the correct amino acids. The growing polypeptide is often termed the nascent chain. Proteins are always biosynthesized from N-terminus to C-terminus.[42]: 1002–42
The size of a synthesized protein can be measured by the number of amino acids it contains and by its total molecular mass, which is normally reported with the unit dalton (Da), or its derivative unit kilodalton (kDa). The average size of a protein increases from Archaea to Bacteria to Eukaryote (283, 311, 438 residues and 31, 34, 49 kDa respectively) due to a bigger number of protein domains constituting proteins in higher organisms.[48] For instance, yeast proteins are on average 466 amino acids long and 53 kDa in mass.[39] The largest known proteins are the titins, a component of the muscle sarcomere, with a molecular mass of almost 3000 kDa and a total length of almost 27000 amino acids.[49]
Chemical synthesis
[edit]
Short proteins can be synthesized chemically by a family of peptide synthesis methods. These rely on organic synthesis techniques such as chemical ligation to produce peptides in high yield.[50] Chemical synthesis allows for the introduction of non-natural amino acids into polypeptide chains, such as attachment of fluorescent probes to amino acid side chains.[51] These methods are useful in laboratory biochemistry and cell biology, though generally not for commercial applications. Chemical synthesis is inefficient for polypeptides longer than about 300 amino acids, and the synthesized proteins may not readily assume their native tertiary structure. Most chemical synthesis methods proceed from C-terminus to N-terminus, opposite the biological reaction.[52]
Structure
[edit]

Most proteins fold into unique 3D structures. The shape into which a protein naturally folds is known as its native conformation.[36]: 36 Although many proteins can fold unassisted, simply through the chemical properties of their amino acids, others require the aid of molecular chaperones to fold into their native states.[36]: 37 Biochemists often refer to four distinct aspects of a protein's structure:[36]: 30–34
- Primary structure: the amino acid sequence. A protein is a polyamide.
- Secondary structure: regularly repeating local structures stabilized by hydrogen bonds. The most common examples are the α-helix, β-sheet and turns. Because secondary structures are local, many regions of distinct secondary structure can be present in the same protein molecule.
- Tertiary structure: the overall shape of a single protein molecule; the spatial relationship of the secondary structures to one another. Tertiary structure is generally stabilized by nonlocal interactions, most commonly the formation of a hydrophobic core, but also through salt bridges, hydrogen bonds, disulfide bonds, and even post-translational modifications. The term "tertiary structure" is often used as synonymous with the term fold. The tertiary structure is what controls the basic function of the protein.
- Quaternary structure: the structure formed by several protein molecules (polypeptide chains), usually called protein subunits in this context, which function as a single protein complex.
- Quinary structure: the signatures of protein surface that organize the crowded cellular interior. Quinary structure is dependent on transient, yet essential, macromolecular interactions that occur inside living cells.
Proteins are not entirely rigid molecules. In addition to these levels of structure, proteins may shift between several related structures while they perform their functions. In the context of these functional rearrangements, these tertiary or quaternary structures are usually referred to as "conformations", and transitions between them are called conformational changes. Such changes are often induced by the binding of a substrate molecule to an enzyme's active site, or the physical region of the protein that participates in chemical catalysis. In solution, protein structures vary because of thermal vibration and collisions with other molecules.[42]: 368–75

Proteins can be informally divided into three main classes, which correlate with typical tertiary structures: globular proteins, fibrous proteins, and membrane proteins. Almost all globular proteins are soluble and many are enzymes. Fibrous proteins are often structural, such as collagen, the major component of connective tissue, or keratin, the protein component of hair and nails. Membrane proteins often serve as receptors or provide channels for polar or charged molecules to pass through the cell membrane.[42]: 165–85
A special case of intramolecular hydrogen bonds within proteins, poorly shielded from water attack and hence promoting their own dehydration, are called dehydrons.[53]
Protein domains
[edit]Many proteins are composed of several protein domains, i.e. segments of a protein that fold into distinct structural units.[54]: 134 Domains usually have specific functions, such as enzymatic activities (e.g. kinase) or they serve as binding modules.[54]: 155–156

Sequence motif
[edit]Short amino acid sequences within proteins often act as recognition sites for other proteins.[56] For instance, SH3 domains typically bind to short PxxP motifs (i.e. 2 prolines [P], separated by two unspecified amino acids [x], although the surrounding amino acids may determine the exact binding specificity). Many such motifs has been collected in the Eukaryotic Linear Motif (ELM) database.[57]
Cellular functions
[edit]Proteins are the chief actors within the cell, said to be carrying out the duties specified by the information encoded in genes.[39] With the exception of certain types of RNA, most other biological molecules are relatively inert elements upon which proteins act. Proteins make up half the dry weight of an Escherichia coli cell, whereas other macromolecules such as DNA and RNA make up only 3% and 20%, respectively.[58] The set of proteins expressed in a particular cell or cell type is known as its proteome.[54]: 120

The chief characteristic of proteins that allows their diverse set of functions is their ability to bind other molecules specifically and tightly. The region of the protein responsible for binding another molecule is known as the binding site and is often a depression or "pocket" on the molecular surface. This binding ability is mediated by the tertiary structure of the protein, which defines the binding site pocket, and by the chemical properties of the surrounding amino acids' side chains. Protein binding can be extraordinarily tight and specific; for example, the ribonuclease inhibitor protein binds to human angiogenin with a sub-femtomolar dissociation constant (< 10−15 M) but does not bind at all to its amphibian homolog onconase (> 1 M). Extremely minor chemical changes such as the addition of a single methyl group to a binding partner can sometimes suffice to nearly eliminate binding; for example, the aminoacyl tRNA synthetase specific to the amino acid valine discriminates against the very similar side chain of the amino acid isoleucine.[59]
Proteins can bind to other proteins as well as to small-molecule substrates. When proteins bind specifically to other copies of the same molecule, they can oligomerize to form fibrils; this process occurs often in structural proteins that consist of globular monomers that self-associate to form rigid fibers. Protein–protein interactions regulate enzymatic activity, control progression through the cell cycle, and allow the assembly of large protein complexes that carry out many closely related reactions with a common biological function. Proteins can bind to, or be integrated into, cell membranes. The ability of binding partners to induce conformational changes in proteins allows the construction of enormously complex signaling networks.[42]: 830–49 As interactions between proteins are reversible and depend heavily on the availability of different groups of partner proteins to form aggregates that are capable to carry out discrete sets of function, study of the interactions between specific proteins is a key to understand important aspects of cellular function, and ultimately the properties that distinguish particular cell types.[60][61]
Enzymes
[edit]The best-known role of proteins in the cell is as enzymes, which catalyse chemical reactions. Enzymes are usually highly specific and accelerate only one or a few chemical reactions. Enzymes carry out most of the reactions involved in metabolism, as well as manipulating DNA in processes such as DNA replication, DNA repair, and transcription. Some enzymes act on other proteins to add or remove chemical groups in a process known as posttranslational modification. About 4,000 reactions are known to be catalysed by enzymes.[62] The rate acceleration conferred by enzymatic catalysis is often enormous—as much as 1017-fold increase in rate over the uncatalysed reaction in the case of orotate decarboxylase (78 million years without the enzyme, 18 milliseconds with the enzyme).[63]
The molecules bound and acted upon by enzymes are called substrates. Although enzymes can consist of hundreds of amino acids, it is usually only a small fraction of the residues that come in contact with the substrate, and an even smaller fraction—three to four residues on average—that are directly involved in catalysis.[64] The region of the enzyme that binds the substrate and contains the catalytic residues is known as the active site.[54]: 389
Dirigent proteins are members of a class of proteins that dictate the stereochemistry of a compound synthesized by other enzymes.[65]
Cell signaling and ligand binding
[edit]
Many proteins are involved in the process of cell signaling and signal transduction. Some proteins, such as insulin, are extracellular proteins that transmit a signal from the cell in which they were synthesized to other cells in distant tissues. Others are membrane proteins that act as receptors whose main function is to bind a signaling molecule and induce a biochemical response in the cell. Many receptors have a binding site exposed on the cell surface and an effector domain within the cell, which may have enzymatic activity or may undergo a conformational change detected by other proteins within the cell.[41]: 251–81
Antibodies are protein components of an adaptive immune system whose main function is to bind antigens, or foreign substances in the body, and target them for destruction. Antibodies can be secreted into the extracellular environment or anchored in the membranes of specialized B cells known as plasma cells. Whereas enzymes are limited in their binding affinity for their substrates by the necessity of conducting their reaction, antibodies have no such constraints. An antibody's binding affinity to its target is extraordinarily high.[42]: 275–50
Many ligand transport proteins bind particular small biomolecules and transport them to other locations in the body of a multicellular organism. These proteins must have a high binding affinity when their ligand is present in high concentrations, and release the ligand when it is present at low concentrations in the target tissues. The canonical example of a ligand-binding protein is haemoglobin, which transports oxygen from the lungs to other organs and tissues in all vertebrates and has close homologs in every biological kingdom.[42]: 222–29 Lectins are sugar-binding proteins which are highly specific for their sugar moieties. Lectins typically play a role in biological recognition phenomena involving cells and proteins.[66] Receptors and hormones are highly specific binding proteins.
Transmembrane proteins can serve as ligand transport proteins that alter the permeability of the cell membrane to small molecules and ions. The membrane alone has a hydrophobic core through which polar or charged molecules cannot diffuse. Membrane proteins contain internal channels that allow such molecules to enter and exit the cell. Many ion channel proteins are specialized to select for only a particular ion; for example, potassium and sodium channels often discriminate for only one of the two ions.[41]: 232–34
Structural proteins
[edit]
Structural proteins confer stiffness and rigidity to otherwise-fluid biological components. Most structural proteins are fibrous proteins; for example, collagen and elastin are critical components of connective tissue such as cartilage, and keratin is found in hard or filamentous structures such as hair, nails, feathers, hooves, and some animal shells.[42]: 178–81 Some globular proteins can play structural functions, for example, actin and tubulin are globular and soluble as monomers, but polymerize to form long, stiff fibers that make up the cytoskeleton, which allows the cell to maintain its shape and size.[54]: 490
Other proteins that serve structural functions are motor proteins such as myosin, kinesin, and dynein, which are capable of generating mechanical forces. These proteins are crucial for cellular motility of single celled organisms and the sperm of many multicellular organisms which reproduce sexually. They generate the forces exerted by contracting muscles[42]: 258–64, 272 and play essential roles in intracellular transport.[54]: 481, 490
Methods of study
[edit]Methods commonly used to study protein structure and function include immunohistochemistry, site-directed mutagenesis, X-ray crystallography, nuclear magnetic resonance and mass spectrometry. The activities and structures of proteins may be examined in vitro, in vivo, and in silico. In vitro studies of purified proteins in controlled environments are useful for learning how a protein carries out its function:[67] for example, enzyme kinetics studies explore the chemical mechanism of an enzyme's catalytic activity and its relative affinity for various possible substrate molecules.[68] By contrast, in vivo experiments can provide information about the physiological role of a protein in the context of a cell or even a whole organism, and can often provide more information about protein behavior in different contexts.[69] In silico studies use computational methods to study proteins.[70]
Protein purification
[edit]Proteins may be purified from other cellular components using a variety of techniques such as ultracentrifugation, precipitation, electrophoresis, and chromatography;[36]: 21–24 the advent of genetic engineering has made possible a number of methods to facilitate purification.[71]
To perform in vitro analysis, a protein must be purified away from other cellular components. This process usually begins with cell lysis, in which a cell's membrane is disrupted and its internal contents released into a solution known as a crude lysate. The resulting mixture can be purified using ultracentrifugation, which fractionates the various cellular components into fractions containing soluble proteins; membrane lipids and proteins; cellular organelles, and nucleic acids. Precipitation by a method known as salting out can concentrate the proteins from this lysate. Various types of chromatography are then used to isolate the protein or proteins of interest based on properties such as molecular weight, net charge and binding affinity.[36]: 21–24 The level of purification can be monitored using various types of gel electrophoresis if the desired protein's molecular weight and isoelectric point are known, by spectroscopy if the protein has distinguishable spectroscopic features, or by enzyme assays if the protein has enzymatic activity. Additionally, proteins can be isolated according to their charge using electrofocusing.[72]
For natural proteins, a series of purification steps may be necessary to obtain protein sufficiently pure for laboratory applications. To simplify this process, genetic engineering is often used to add chemical features to proteins that make them easier to purify without affecting their structure or activity. Here, a "tag" consisting of a specific amino acid sequence, often a series of histidine residues (a "His-tag"), is attached to one terminus of the protein. As a result, when the lysate is passed over a chromatography column containing nickel, the histidine residues ligate the nickel and attach to the column while the untagged components of the lysate pass unimpeded. A number of tags have been developed to help researchers purify specific proteins from complex mixtures.[71]
Cellular localization
[edit]
The study of proteins in vivo is often concerned with the synthesis and localization of the protein within the cell. Although many intracellular proteins are synthesized in the cytoplasm and membrane-bound or secreted proteins in the endoplasmic reticulum, the specifics of how proteins are targeted to specific organelles or cellular structures is often unclear. A useful technique for assessing cellular localization uses genetic engineering to express in a cell a fusion protein or chimera consisting of the natural protein of interest linked to a "reporter" such as green fluorescent protein (GFP).[73] The fused protein's position within the cell can then be cleanly and efficiently visualized using microscopy.[74]
Other methods for elucidating the cellular location of proteins requires the use of known compartmental markers for regions such as the ER, the Golgi, lysosomes or vacuoles, mitochondria, chloroplasts, plasma membrane, etc. With the use of fluorescently tagged versions of these markers or of antibodies to known markers, it becomes much simpler to identify the localization of a protein of interest. For example, indirect immunofluorescence will allow for fluorescence colocalization and demonstration of location. Fluorescent dyes are used to label cellular compartments for a similar purpose.[75]
Other possibilities exist, as well. For example, immunohistochemistry usually uses an antibody to one or more proteins of interest that are conjugated to enzymes yielding either luminescent or chromogenic signals that can be compared between samples, allowing for localization information.[76] Another applicable technique is cofractionation in sucrose (or other material) gradients using isopycnic centrifugation.[77] While this technique does not prove colocalization of a compartment of known density and the protein of interest, it indicates an increased likelihood.[77]
Finally, the gold-standard method of cellular localization is immunoelectron microscopy. This technique uses an antibody to the protein of interest, along with classical electron microscopy techniques. The sample is prepared for normal electron microscopic examination, and then treated with an antibody to the protein of interest that is conjugated to an extremely electro-dense material, usually gold. This allows for the localization of both ultrastructural details as well as the protein of interest.[78]
Through another genetic engineering application known as site-directed mutagenesis, researchers can alter the protein sequence and hence its structure, cellular localization, and susceptibility to regulation. This technique even allows the incorporation of unnatural amino acids into proteins, using modified tRNAs,[79] and may allow the rational design of new proteins with novel properties.[80]
Proteomics
[edit]The total complement of proteins present at a time in a cell or cell type is known as its proteome, and the study of such large-scale data sets defines the field of proteomics, named by analogy to the related field of genomics. Key experimental techniques in proteomics include 2D electrophoresis,[81] which allows the separation of many proteins, mass spectrometry,[82] which allows rapid high-throughput identification of proteins and sequencing of peptides (most often after in-gel digestion), protein microarrays, which allow the detection of the relative levels of the various proteins present in a cell, and two-hybrid screening, which allows the systematic exploration of protein–protein interactions.[83] The total complement of biologically possible such interactions is known as the interactome.[84] A systematic attempt to determine the structures of proteins representing every possible fold is known as structural genomics.[85]
Structure determination
[edit]Discovering the tertiary structure of a protein, or the quaternary structure of its complexes, can provide important clues about how the protein performs its function and how it can be affected, i.e. in drug design. As proteins are too small to be seen under a light microscope, other methods have to be employed to determine their structure. Common experimental methods include X-ray crystallography and NMR spectroscopy, both of which can produce structural information at atomic resolution. However, NMR experiments are able to provide information from which a subset of distances between pairs of atoms can be estimated, and the final possible conformations for a protein are determined by solving a distance geometry problem. Dual polarisation interferometry is a quantitative analytical method for measuring the overall protein conformation and conformational changes due to interactions or other stimulus. Circular dichroism is another laboratory technique for determining internal β-sheet / α-helical composition of proteins. Cryoelectron microscopy is used to produce lower-resolution structural information about very large protein complexes, including assembled viruses;[41]: 340–41 a variant known as electron crystallography can produce high-resolution information in some cases, especially for two-dimensional crystals of membrane proteins.[86] Solved structures are usually deposited in the Protein Data Bank (PDB), a freely available resource from which structural data about thousands of proteins can be obtained in the form of Cartesian coordinates for each atom in the protein.[87]
Many more gene sequences are known than protein structures. Further, the set of solved structures is biased toward proteins that can be easily subjected to the conditions required in X-ray crystallography, one of the major structure determination methods. In particular, globular proteins are comparatively easy to crystallize in preparation for X-ray crystallography. Membrane proteins and large protein complexes, by contrast, are difficult to crystallize and are underrepresented in the PDB.[88] Structural genomics initiatives have attempted to remedy these deficiencies by systematically solving representative structures of major fold classes. Protein structure prediction methods attempt to provide a means of generating a plausible structure for proteins whose structures have not been experimentally determined.[89]
Structure prediction
[edit]
Complementary to the field of structural genomics, protein structure prediction develops efficient mathematical models of proteins to computationally predict the molecular formations in theory, instead of detecting structures with laboratory observation.[90] The most successful type of structure prediction, known as homology modeling, relies on the existence of a "template" structure with sequence similarity to the protein being modeled; structural genomics' goal is to provide sufficient representation in solved structures to model most of those that remain.[91] Although producing accurate models remains a challenge when only distantly related template structures are available, it has been suggested that sequence alignment is the bottleneck in this process, as quite accurate models can be produced if a "perfect" sequence alignment is known.[92] Many structure prediction methods have served to inform the emerging field of protein engineering, in which novel protein folds have already been designed.[93] Many proteins (in eukaryotes ~33%) contain large unstructured but biologically functional segments and can be classified as intrinsically disordered proteins. Predicting and analysing protein disorder is an important part of protein structure characterisation.[94]
In silico simulation of dynamical processes
[edit]A more complex computational problem is the prediction of intermolecular interactions, such as in molecular docking,[95] protein folding, protein–protein interaction and chemical reactivity. Mathematical models to simulate these dynamical processes involve molecular mechanics, in particular, molecular dynamics. In this regard, in silico simulations discovered the folding of small α-helical protein domains such as the villin headpiece,[96] the HIV accessory protein[97] and hybrid methods combining standard molecular dynamics with quantum mechanical mathematics have explored the electronic states of rhodopsins.[98]
Beyond classical molecular dynamics, quantum dynamics methods allow the simulation of proteins in atomistic detail with an accurate description of quantum mechanical effects. Examples include the multi-layer multi-configuration time-dependent Hartree method and the hierarchical equations of motion approach, which have been applied to plant cryptochromes[99] and bacteria light-harvesting complexes,[100] respectively. Both quantum and classical mechanical simulations of biological-scale systems are extremely computationally demanding, so distributed computing initiatives such as the Folding@home project facilitate the molecular modeling by exploiting advances in GPU parallel processing and Monte Carlo techniques.[101][102]
Chemical analysis
[edit]The total nitrogen content of organic matter is mainly formed by the amino groups in proteins. The total Kjeldahl nitrogen (TKN) is a measure of nitrogen widely used in the analysis of (waste) water, soil, food, feed and organic matter in general. As the name suggests, the Kjeldahl method is applied. More sensitive methods are available.[103][104]
Digestion
[edit]
In the absence of catalysts, proteins are slow to hydrolyze.[105] The breakdown of proteins to small peptides and amino acids (proteolysis) is a step in digestion; these breakdown products are then absorbed in the small intestine.[106] The hydrolysis of proteins relies on enzymes called proteases or peptidases. Proteases, which are themselves proteins, come in several types according to the particular peptide bonds that they cleave as well as their tendency to cleave peptide bonds at the terminus of a protein (exopeptidases) vs peptide bonds at the interior of the protein (endopeptidases).[107] Pepsin is an endopeptidase in the stomach. Subsequent to the stomach, the pancreas secretes other proteases to complete the hydrolysis, these include trypsin and chymotrypsin.[108]
Protein hydrolysis is employed commercially as a means of producing amino acids from bulk sources of protein, such as blood meal, feathers, keratin. Such materials are treated with hot hydrochloric acid, which effects the hydrolysis of the peptide bonds.[109]
Mechanical properties
[edit]The mechanical properties of proteins are highly diverse and are often central to their biological function, as in the case of proteins like keratin and collagen.[110] For instance, the ability of muscle tissue to continually expand and contract is directly tied to the elastic properties of their underlying protein makeup.[111][112] Beyond fibrous proteins, the conformational dynamics of enzymes[113] and the structure of biological membranes, among other biological functions, are governed by the mechanical properties of the proteins. Outside of their biological context, the unique mechanical properties of many proteins, along with their relative sustainability when compared to synthetic polymers, have made them desirable targets for next-generation materials design.[114][115]
Young's modulus, E, is calculated as the axial stress σ over the resulting strain ε. It is a measure of the relative stiffness of a material. In the context of proteins, this stiffness often directly correlates to biological function. For example, collagen, found in connective tissue, bones, and cartilage, and keratin, found in nails, claws, and hair, have observed stiffnesses that are several orders of magnitude higher than that of elastin,[116] which is though to give elasticity to structures such as blood vessels, pulmonary tissue, and bladder tissue, among others.[117][118] In comparison to this, globular proteins, such as Bovine Serum Albumin, which float relatively freely in the cytosol and often function as enzymes (and thus undergoing frequent conformational changes) have comparably much lower Young's moduli.[119][120]
The Young's modulus of a single protein can be found through molecular dynamics simulation. Using either atomistic force-fields, such as CHARMM or GROMOS, or coarse-grained forcefields like Martini,[121] a single protein molecule can be stretched by a uniaxial force while the resulting extension is recorded in order to calculate the strain.[122][123] Experimentally, methods such as atomic force microscopy can be used to obtain similar data.[124] The internal dynamics of proteins involve subtle elastic and plastic deformations induced by viscoelastic forces, which can be probed by nano-rheology techniques.[125] These estimates yield typical spring constants around k ≈ 100 pN/nm, equivalent to Young's moduli of E ≈ 100 MPa, and typical friction coefficients of γ ≈ 0.1 pN·s/nm, corresponding to viscosity of η ≈ 0.01 pN·s/nm2 = 107 cP (that is, 107 times more viscous than water).
At the macroscopic level, the Young's modulus of cross-linked protein networks can be obtained through more traditional mechanical testing. Experimentally observed values for a few proteins can be seen below.
| Protein | Protein class | Young's modulus |
|---|---|---|
| keratin (cross-linked) | fibrous | 1.5–10 GPa[126] |
| elastin (cross-linked) | fibrous | 1 MPa[116] |
| fibrin (cross-linked) | fibrous | 1–10 MPa[116] |
| collagen (cross-linked) | fibrous | 5–7.5 GPa[116][127] |
| resilin (cross-linked) | fibrous | 1–2 MPa[116] |
| bovine serum albumin (cross-linked) | globular | 2.5–15 kPa[119] |
| β-barrel outer membrane proteins | membrane | 20–45 GPa[128] |
See also
[edit]References
[edit]- ^ a b c Osborne TB (1909). "History". The Vegetable Proteins. pp. 1–6.
- ^ a b c Reynolds JA, Tanford C (2003). Nature's Robots: A History of Proteins (Oxford Paperbacks). New York, New York: Oxford University Press. p. 15. ISBN 978-0-19-860694-9.
- ^ Tanford C (2001). Nature's robots: a history of proteins. Internet Archive. Oxford; Toronto: Oxford University Press. ISBN 978-0-19-850466-5.
- ^ Mulder GJ (1838). "Sur la composition de quelques substances animales". Bulletin des Sciences Physiques et Naturelles en Néerlande: 104.
- ^ Hartley H (August 1951). "Origin of the word 'protein'". Nature. 168 (4267): 244. Bibcode:1951Natur.168..244H. doi:10.1038/168244a0. PMID 14875059. S2CID 4271525.
- ^ a b c Perrett D (August 2007). "From 'protein' to the beginnings of clinical proteomics". Proteomics. Clinical Applications. 1 (8): 720–738. doi:10.1002/prca.200700525. PMID 21136729. S2CID 32843102.
- ^ "Protein (n.)". Oxford English Dictionary. July 2023. doi:10.1093/OED/5657543824.
- ^ Bischoff TL, Voit C (1860). Die Gesetze der Ernaehrung des Pflanzenfressers durch neue Untersuchungen festgestellt (in German). Leipzig, Heidelberg: C.F. Winter'sche Verlagshandlung.
- ^ Osborne TB (April 1913). "In Memoriam Heinrich Ritthausen". Biochemical Bulletin. II (7). Columbia University Biochemical Association: 338. Retrieved 1 January 2016., archived at the Biodiversity Heritage Library
- ^ Simoni RD, Hill RL, Vaughan M (2002-05-03). "Nutritional Biochemistry and the Amino Acid Composition of Proteins: The early years of protein chemistry, the Work of Thomas B. Osborne and Lafayette B. Mendel". Journal of Biological Chemistry. 277 (18): 14–15. doi:10.1016/S0021-9258(19)35800-4.
- ^ Osborne TB, Mendel LB, Ferry EL, Wakeman AJ (1916). "The Amino-Acid Minimum for Maintenance and Growth, as Exemplified by Further Experiments with Lysine and Tryptophane". Journal of Biological Chemistry. 25 (1): 1–12. doi:10.1016/S0021-9258(18)87509-3.
- ^ Simoni RD, Hill RL, Vaughan M (2002-09-13). "The Discovery of the Amino Acid Threonine: the Work of William C. Rose". Journal of Biological Chemistry. 277 (37): 56–58. doi:10.1016/S0021-9258(20)74369-3.
- ^ "Hofmeister, Franz". encyclopedia.com. Archived from the original on 5 April 2017. Retrieved 4 April 2017.
- ^ Koshland DE, Haurowitz F. "Protein, section: Classification of protein". britannica.com. Archived from the original on 4 April 2017. Retrieved 4 April 2017.
- ^ Sumner JB (August 1926). "The Isolation and Crystallization of the Enzyme Urease". Journal of Biological Chemistry. 69 (2): 435–441. doi:10.1016/S0021-9258(18)84560-4.
- ^ Pauling L, Corey RB (May 1951). "Atomic coordinates and structure factors for two helical configurations of polypeptide chains". Proceedings of the National Academy of Sciences of the United States of America. 37 (5): 235–240. Bibcode:1951PNAS...37..235P. doi:10.1073/pnas.37.5.235. PMC 1063348. PMID 14834145.
- ^ Kauzmann W (May 1956). "Structural factors in protein denaturation". Journal of Cellular Physiology. Supplement. 47 (Suppl 1): 113–131. doi:10.1002/jcp.1030470410. PMID 13332017.
- ^ Kauzmann W (1959) [1959]. "Some Factors in the Interpretation of Protein Denaturation". In Anfinsen Jr C, Anson M, Bailey K, Edsall JT (eds.). Advances in Protein Chemistry. Vol. 14. Elsevier. pp. 1–63. doi:10.1016/s0065-3233(08)60608-7. ISBN 978-0-12-034214-3. PMID 14404936. Retrieved 2025-04-21.
{{cite book}}: ISBN / Date incompatibility (help) - ^ Kalman SM, Linderstrøm-Lang K, Ottesen M, Richards FM (February 1955). "Degradation of ribonuclease by subtilisin". Biochimica et Biophysica Acta. 16 (2): 297–299. doi:10.1016/0006-3002(55)90224-9. PMID 14363272.
- ^ Dill KA (1990). "Dominant forces in protein folding". Biochemistry. 29 (31): 7133–7155. doi:10.1021/bi00483a001. PMID 2207096.
- ^ Sanger F (1949). "The terminal peptides of insulin". The Biochemical Journal. 45 (5): 563–574. doi:10.1042/bj0450563. PMC 1275055. PMID 15396627.
- ^ Sanger F. (1958), Nobel lecture: The chemistry of insulin (PDF), Nobelprize.org, archived (PDF) from the original on 2013-03-19, retrieved 2016-02-09
- ^ Richards FM (1972). "The 1972 nobel prize for chemistry". Science. 178 (4060): 492–3. Bibcode:1972Sci...178..492R. doi:10.1126/science.178.4060.492. PMID 17754377.
- ^ Marshall GR, Feng JA, Kuster DJ (2008). "Back to the future: Ribonuclease A". Biopolymers. 90 (3): 259–77. doi:10.1002/bip.20845. PMID 17868092.
- ^ a b c Stoddart C (1 March 2022). "Structural biology: How proteins got their close-up". Knowable Magazine. doi:10.1146/knowable-022822-1.
- ^ Muirhead H, Perutz MF (August 1963). ". Structure of haemoglobin. A three-dimensional Fourier synthesis of reduced human haemoglobin at 5.5 Å resolution". Nature. 199 (4894): 633–638. Bibcode:1963Natur.199..633M. doi:10.1038/199633a0. PMID 14074546. S2CID 4257461.
- ^ Kendrew JC, Bodo G, Dintzis HM, Parrish RG, Wyckoff H, Phillips DC (March 1958). "A three-dimensional model of the myoglobin molecule obtained by x-ray analysis". Nature. 181 (4610): 662–666. Bibcode:1958Natur.181..662K. doi:10.1038/181662a0. PMID 13517261. S2CID 4162786.
- ^ Zhou ZH (April 2008). "Towards atomic resolution structural determination by single-particle cryo-electron microscopy". Current Opinion in Structural Biology. 18 (2): 218–228. doi:10.1016/j.sbi.2008.03.004. PMC 2714865. PMID 18403197.
- ^ Keskin O, Tuncbag N, Gursoy A (April 2008). "Characterization and prediction of protein interfaces to infer protein-protein interaction networks". Current Pharmaceutical Biotechnology. 9 (2): 67–76. doi:10.2174/138920108783955191. hdl:11511/32640. PMID 18393863.
- ^ "Summary Statistics". RCSB PDB. Retrieved 2024-04-20.
- ^ McDonald AG, Tipton KF (2023). "Enzyme nomenclature and classification: the state of the art". The FEBS Journal. 290 (9): 2214–2231. doi:10.1111/febs.16274. ISSN 1742-464X. PMID 34773359.
- ^ The Gene Ontology Consortium (January 2008). "The Gene Ontology project in 2008". Nucleic Acids Research. 36 (Database issue): D440–4. doi:10.1093/nar/gkm883. PMC 2238979. PMID 17984083.
- ^ Ekman D, Björklund AK, Frey-Skött J, Elofsson A (April 2005). "Multi-domain proteins in the three kingdoms of life: orphan domains and other unassigned regions". Journal of Molecular Biology. 348 (1): 231–243. doi:10.1016/j.jmb.2005.02.007. PMID 15808866.
- ^ Nelson DL, Cox MM (2005). Lehninger's Principles of Biochemistry (4th ed.). New York, New York: W. H. Freeman and Company. Bibcode:2005lpbc.book.....N.
- ^ Gutteridge A, Thornton JM (November 2005). "Understanding nature's catalytic toolkit". Trends in Biochemical Sciences. 30 (11): 622–629. doi:10.1016/j.tibs.2005.09.006. PMID 16214343.
- ^ a b c d e f g Murray RF, Harper HW, Granner DK, Mayes PA, Rodwell VW (2006). Harper's Illustrated Biochemistry. New York: Lange Medical Books/McGraw-Hill. ISBN 978-0-07-146197-9.
- ^ a b Reusch W (5 May 2013). "Peptides & Proteins". Michigan State University Department of Chemistry.
- ^ Stryer L (2002). Biochemistry (Fifth ed.). W. H. Freeman and Company. p. 826. ISBN 0-7167-4684-0.
{{cite book}}: CS1 maint: overridden setting (link) - ^ a b c Lodish H, Berk A, Matsudaira P, Kaiser CA, Krieger M, Scott MP, et al. (2004). Molecular Cell Biology (5th ed.). New York, New York: WH Freeman and Company.
- ^ Ardejani MS, Powers ET, Kelly JW (August 2017). "Using Cooperatively Folded Peptides To Measure Interaction Energies and Conformational Propensities". Accounts of Chemical Research. 50 (8): 1875–1882. doi:10.1021/acs.accounts.7b00195. PMC 5584629. PMID 28723063.
- ^ a b c d Branden C, Tooze J (1999). Introduction to Protein Structure. New York: Garland Pub. ISBN 978-0-8153-2305-1.
- ^ a b c d e f g h i j Van Holde KE, Mathews CK (1996). Biochemistry. Menlo Park, California: Benjamin/Cummings. ISBN 978-0-8053-3931-4.
- ^ Milo R (December 2013). "What is the total number of protein molecules per cell volume? A call to rethink some published values". BioEssays. 35 (12): 1050–1055. doi:10.1002/bies.201300066. PMC 3910158. PMID 24114984.
- ^ Beck M, Schmidt A, Malmstroem J, Claassen M, Ori A, Szymborska A, et al. (November 2011). "The quantitative proteome of a human cell line". Molecular Systems Biology. 7: 549. doi:10.1038/msb.2011.82. PMC 3261713. PMID 22068332.
- ^ Wu L, Candille SI, Choi Y, Xie D, Jiang L, Li-Pook-Than J, et al. (July 2013). "Variation and genetic control of protein abundance in humans". Nature. 499 (7456): 79–82. Bibcode:2013Natur.499...79W. doi:10.1038/nature12223. PMC 3789121. PMID 23676674.
- ^ Ellis R (1979). "The most abundant protein in the world". Trends in Biochemical Sciences. 4 (11): 241–244. doi:10.1016/0968-0004(79)90212-3.
- ^ Dobson CM (2000). "The nature and significance of protein folding". In Pain RH (ed.). Mechanisms of Protein Folding. Oxford, Oxfordshire: Oxford University Press. pp. 1–28. ISBN 978-0-19-963789-8.
- ^ Kozlowski LP (January 2017). "Proteome-pI: proteome isoelectric point database". Nucleic Acids Research. 45 (D1): D1112 – D1116. doi:10.1093/nar/gkw978. PMC 5210655. PMID 27789699.
- ^ Fulton AB, Isaacs WB (April 1991). "Titin, a huge, elastic sarcomeric protein with a probable role in morphogenesis". BioEssays. 13 (4): 157–161. doi:10.1002/bies.950130403. PMID 1859393. S2CID 20237314.
- ^ Bruckdorfer T, Marder O, Albericio F (February 2004). "From production of peptides in milligram amounts for research to multi-tons quantities for drugs of the future". Current Pharmaceutical Biotechnology. 5 (1): 29–43. doi:10.2174/1389201043489620. PMID 14965208.
- ^ Schwarzer D, Cole PA (December 2005). "Protein semisynthesis and expressed protein ligation: chasing a protein's tail". Current Opinion in Chemical Biology. 9 (6): 561–569. doi:10.1016/j.cbpa.2005.09.018. PMID 16226484.
- ^ Kent SB (February 2009). "Total chemical synthesis of proteins". Chemical Society Reviews. 38 (2): 338–351. doi:10.1039/b700141j. PMID 19169452. S2CID 5432012.
- ^ Fernández A, Scott R (September 2003). "Dehydron: a structurally encoded signal for protein interaction". Biophysical Journal. 85 (3): 1914–1928. Bibcode:2003BpJ....85.1914F. doi:10.1016/S0006-3495(03)74619-0. PMC 1303363. PMID 12944304.
- ^ a b c d e f Garrett R, Grisham CM (2010). Biochemistry (4th ed.). Belmont, CA: Brooks/Cole, Cengage Learning. ISBN 978-0-495-10935-8.
- ^ Drees F, Gertler FB (2008-02-01). "Ena/VASP: proteins at the tip of the nervous system". Current Opinion in Neurobiology. Development. 18 (1): 53–59. doi:10.1016/j.conb.2008.05.007. PMC 2515615. PMID 18508258.
- ^ Davey NE, Van Roey K, Weatheritt RJ, Toedt G, Uyar B, Altenberg B, et al. (January 2012). "Attributes of short linear motifs". Molecular BioSystems. 8 (1): 268–281. doi:10.1039/c1mb05231d. PMID 21909575.
- ^ Kumar M, Gouw M, Michael S, Sámano-Sánchez H, Pancsa R, Glavina J, et al. (2020-01-08). "ELM-the eukaryotic linear motif resource in 2020". Nucleic Acids Research. 48 (D1): D296 – D306. doi:10.1093/nar/gkz1030. PMC 7145657. PMID 31680160.
- ^ Voet D, Voet JG. (2004). Biochemistry Vol 1 3rd ed. Wiley: Hoboken, NJ.
- ^ Sankaranarayanan R, Moras D (2001). "The fidelity of the translation of the genetic code". Acta Biochimica Polonica. 48 (2): 323–335. doi:10.18388/abp.2001_3918. PMID 11732604.
- ^ Copland JA, Sheffield-Moore M, Koldzic-Zivanovic N, Gentry S, Lamprou G, Tzortzatou-Stathopoulou F, et al. (June 2009). "Sex steroid receptors in skeletal differentiation and epithelial neoplasia: is tissue-specific intervention possible?". BioEssays. 31 (6): 629–641. doi:10.1002/bies.200800138. PMID 19382224. S2CID 205469320.
- ^ Samarin S, Nusrat A (January 2009). "Regulation of epithelial apical junctional complex by Rho family GTPases". Frontiers in Bioscience. 14 (3): 1129–1142. doi:10.2741/3298. PMID 19273120.
- ^ Bairoch A (January 2000). "The ENZYME database in 2000". Nucleic Acids Research. 28 (1): 304–305. doi:10.1093/nar/28.1.304. PMC 102465. PMID 10592255.
- ^ Radzicka A, Wolfenden R (January 1995). "A proficient enzyme". Science. 267 (5194): 90–93. Bibcode:1995Sci...267...90R. doi:10.1126/science.7809611. PMID 7809611.
- ^ EBI External Services (2010-01-20). "The Catalytic Site Atlas at The European Bioinformatics Institute". Ebi.ac.uk. Archived from the original on 2013-08-03. Retrieved 2011-01-16.
- ^ Pickel B, Schaller A (October 2013). "Dirigent proteins: molecular characteristics and potential biotechnological applications". Applied Microbiology and Biotechnology. 97 (19): 8427–8438. doi:10.1007/s00253-013-5167-4. PMID 23989917. S2CID 1896003.
- ^ Rüdiger H, Siebert HC, Solís D, Jiménez-Barbero J, Romero A, von der Lieth CW, et al. (April 2000). "Medicinal chemistry based on the sugar code: fundamentals of lectinology and experimental strategies with lectins as targets". Current Medicinal Chemistry. 7 (4): 389–416. doi:10.2174/0929867003375164. PMID 10702616.
- ^ National Research Council (US) Subcommittee on Reproductive and Developmental Toxicity (2001), "Experimental Animal and In Vitro Study Designs", Evaluating Chemical and Other Agent Exposures for Reproductive and Developmental Toxicity, National Academies Press, retrieved 2024-12-23
- ^ Ricard J, Cornish-Bowden A (July 1987). "Co-operative and allosteric enzymes: 20 years on". European Journal of Biochemistry. 166 (2): 255–272. doi:10.1111/j.1432-1033.1987.tb13510.x. PMID 3301336.
- ^ Lipinski C, Hopkins A (December 2004). "Navigating chemical space for biology and medicine". Nature. 432 (7019): 855–861. Bibcode:2004Natur.432..855L. doi:10.1038/nature03193. PMID 15602551.
- ^ Danchin A, Médigue C, Gascuel O, Soldano H, Hénaut A (1991). "From data banks to data bases". Research in Microbiology. 142 (7–8): 913–916. doi:10.1016/0923-2508(91)90073-J. PMID 1784830.
- ^ a b Terpe K (January 2003). "Overview of tag protein fusions: from molecular and biochemical fundamentals to commercial systems". Applied Microbiology and Biotechnology. 60 (5): 523–533. doi:10.1007/s00253-002-1158-6. PMID 12536251. S2CID 206934268.
- ^ Hey J, Posch A, Cohen A, Liu N, Harbers A (2008). "Fractionation of Complex Protein Mixtures by Liquid-Phase Isoelectric Focusing". 2D PAGE: Sample Preparation and Fractionation. Methods in Molecular Biology. Vol. 424. pp. 225–239. doi:10.1007/978-1-60327-064-9_19. ISBN 978-1-58829-722-8. PMID 18369866.
- ^ Stepanenko OV, Verkhusha VV, Kuznetsova IM, Uversky VN, Turoverov KK (August 2008). "Fluorescent proteins as biomarkers and biosensors: throwing color lights on molecular and cellular processes". Current Protein & Peptide Science. 9 (4): 338–369. doi:10.2174/138920308785132668. PMC 2904242. PMID 18691124.
- ^ Yuste R (December 2005). "Fluorescence microscopy today". Nature Methods. 2 (12): 902–904. doi:10.1038/nmeth1205-902. PMID 16299474. S2CID 205418407.
- ^ Margolin W (January 2000). "Green fluorescent protein as a reporter for macromolecular localization in bacterial cells". Methods. 20 (1): 62–72. doi:10.1006/meth.1999.0906. PMID 10610805.
- ^ Hrycaj S (17 October 2023). "Immunohistochemistry: Origins, Tips, and a Look to the Future". The Scientist Magazine. Retrieved 2024-12-22.
- ^ a b Walker JH, Wilson K (2000). Principles and Techniques of Practical Biochemistry. Cambridge, UK: Cambridge University Press. pp. 287–89. ISBN 978-0-521-65873-7.
- ^ Mayhew TM, Lucocq JM (August 2008). "Developments in cell biology for quantitative immunoelectron microscopy based on thin sections: a review". Histochemistry and Cell Biology. 130 (2) 299: 299–313. doi:10.1007/s00418-008-0451-6. PMC 2491712. PMID 18553098.
- ^ Hohsaka T, Sisido M (December 2002). "Incorporation of non-natural amino acids into proteins". Current Opinion in Chemical Biology. 6 (6): 809–815. doi:10.1016/S1367-5931(02)00376-9. PMID 12470735.
- ^ Cedrone F, Ménez A, Quéméneur E (August 2000). "Tailoring new enzyme functions by rational redesign". Current Opinion in Structural Biology. 10 (4): 405–410. doi:10.1016/S0959-440X(00)00106-8. PMID 10981626.
- ^ Görg A, Weiss W, Dunn MJ (December 2004). "Current two-dimensional electrophoresis technology for proteomics". Proteomics. 4 (12): 3665–3685. doi:10.1002/pmic.200401031. PMID 15543535. S2CID 28594824.
- ^ Conrotto P, Souchelnytskyi S (September 2008). "Proteomic approaches in biological and medical sciences: principles and applications". Experimental Oncology. 30 (3): 171–180. PMID 18806738.
- ^ Koegl M, Uetz P (December 2007). "Improving yeast two-hybrid screening systems". Briefings in Functional Genomics & Proteomics. 6 (4): 302–312. doi:10.1093/bfgp/elm035. PMID 18218650.
- ^ Plewczyński D, Ginalski K (2009). "The interactome: predicting the protein-protein interactions in cells". Cellular & Molecular Biology Letters. 14 (1): 1–22. doi:10.2478/s11658-008-0024-7. PMC 6275871. PMID 18839074.
- ^ Zhang C, Kim SH (February 2003). "Overview of structural genomics: from structure to function". Current Opinion in Chemical Biology. 7 (1): 28–32. doi:10.1016/S1367-5931(02)00015-7. PMID 12547423.
- ^ Gonen T, Cheng Y, Sliz P, Hiroaki Y, Fujiyoshi Y, Harrison SC, et al. (December 2005). "Lipid-protein interactions in double-layered two-dimensional AQP0 crystals". Nature. 438 (7068): 633–638. Bibcode:2005Natur.438..633G. doi:10.1038/nature04321. PMC 1350984. PMID 16319884.
- ^ Standley DM, Kinjo AR, Kinoshita K, Nakamura H (July 2008). "Protein structure databases with new web services for structural biology and biomedical research". Briefings in Bioinformatics. 9 (4): 276–285. doi:10.1093/bib/bbn015. PMID 18430752.
- ^ Walian P, Cross TA, Jap BK (2004). "Structural genomics of membrane proteins". Genome Biology. 5 (4): 215. doi:10.1186/gb-2004-5-4-215. PMC 395774. PMID 15059248.
- ^ Sleator RD (2012). "Prediction of Protein Functions". Functional Genomics. Methods in Molecular Biology. Vol. 815. pp. 15–24. doi:10.1007/978-1-61779-424-7_2. ISBN 978-1-61779-423-0. PMID 22130980.
- ^ Zhang Y (June 2008). "Progress and challenges in protein structure prediction". Current Opinion in Structural Biology. 18 (3): 342–348. doi:10.1016/j.sbi.2008.02.004. PMC 2680823. PMID 18436442.
- ^ Xiang Z (June 2006). "Advances in homology protein structure modeling". Current Protein & Peptide Science. 7 (3): 217–227. doi:10.2174/138920306777452312. PMC 1839925. PMID 16787261.
- ^ Zhang Y, Skolnick J (January 2005). "The protein structure prediction problem could be solved using the current PDB library". Proceedings of the National Academy of Sciences of the United States of America. 102 (4): 1029–1034. Bibcode:2005PNAS..102.1029Z. doi:10.1073/pnas.0407152101. PMC 545829. PMID 15653774.
- ^ Kuhlman B, Dantas G, Ireton GC, Varani G, Stoddard BL, Baker D (November 2003). "Design of a novel globular protein fold with atomic-level accuracy". Science. 302 (5649): 1364–1368. Bibcode:2003Sci...302.1364K. doi:10.1126/science.1089427. PMID 14631033. S2CID 1939390.
- ^ Ward JJ, Sodhi JS, McGuffin LJ, Buxton BF, Jones DT (March 2004). "Prediction and functional analysis of native disorder in proteins from the three kingdoms of life". Journal of Molecular Biology. 337 (3): 635–645. CiteSeerX 10.1.1.120.5605. doi:10.1016/j.jmb.2004.02.002. PMID 15019783.
- ^ Ritchie DW (February 2008). "Recent progress and future directions in protein-protein docking". Current Protein & Peptide Science. 9 (1): 1–15. CiteSeerX 10.1.1.211.4946. doi:10.2174/138920308783565741. PMID 18336319.
- ^ Zagrovic B, Snow CD, Shirts MR, Pande VS (November 2002). "Simulation of folding of a small alpha-helical protein in atomistic detail using worldwide-distributed computing". Journal of Molecular Biology. 323 (5): 927–937. CiteSeerX 10.1.1.142.8664. doi:10.1016/S0022-2836(02)00997-X. PMID 12417204.
- ^ Herges T, Wenzel W (January 2005). "In silico folding of a three helix protein and characterization of its free-energy landscape in an all-atom force field". Physical Review Letters. 94 (1) 018101. arXiv:physics/0310146. Bibcode:2005PhRvL..94a8101H. doi:10.1103/PhysRevLett.94.018101. PMID 15698135. S2CID 1477100.
- ^ Hoffmann M, Wanko M, Strodel P, König PH, Frauenheim T, Schulten K, et al. (August 2006). "Color tuning in rhodopsins: the mechanism for the spectral shift between bacteriorhodopsin and sensory rhodopsin II". Journal of the American Chemical Society. 128 (33): 10808–10818. Bibcode:2006JAChS.12810808H. doi:10.1021/ja062082i. PMID 16910676.
- ^ Mendive-Tapia D, Mangaud E, Firmino T, de la Lande A, Desouter-Lecomte M, Meyer HD, et al. (January 2018). "Multidimensional Quantum Mechanical Modeling of Electron Transfer and Electronic Coherence in Plant Cryptochromes: The Role of Initial Bath Conditions". The Journal of Physical Chemistry B. 122 (1): 126–136. doi:10.1021/acs.jpcb.7b10412. PMID 29216421.
- ^ Strümpfer J, Schulten K (August 2012). "Open Quantum Dynamics Calculations with the Hierarchy Equations of Motion on Parallel Computers". Journal of Chemical Theory and Computation. 8 (8): 2808–2816. doi:10.1021/ct3003833. PMC 3480185. PMID 23105920.
- ^ Scheraga HA, Khalili M, Liwo A (2007). "Protein-folding dynamics: overview of molecular simulation techniques". Annual Review of Physical Chemistry. 58: 57–83. Bibcode:2007ARPC...58...57S. doi:10.1146/annurev.physchem.58.032806.104614. PMID 17034338.
- ^ Zheng S, Javidpour L, Sahimi M, Shing KS, Nakano A (2020). "sDMD: An open source program for discontinuous molecular dynamics simulation of protein folding and aggregation". Computer Physics Communications. 247 106873. Bibcode:2020CoPhC.24706873Z. doi:10.1016/j.cpc.2019.106873.
- ^ Muñoz-Huerta RF, Guevara-Gonzalez RG, Contreras-Medina LM, Torres-Pacheco I, Prado-Olivarez J, Ocampo-Velazquez RV (August 2013). "A review of methods for sensing the nitrogen status in plants: advantages, disadvantages and recent advances". Sensors. 13 (8): 10823–10843. Bibcode:2013Senso..1310823M. doi:10.3390/s130810823. PMC 3812630. PMID 23959242.
- ^ Martin PD, Malley DF, Manning G, Fuller L (November 2002). "Determination of soil organic carbon and nitrogen at the field level using near-infrared spectroscopy". Canadian Journal of Soil Science. 82 (4): 413–422. Bibcode:2002CaJSS..82..413M. doi:10.4141/S01-054.
- ^ Radzicka A, Wolfenden R (1 January 1996). "Rates of Uncatalyzed Peptide Bond Hydrolysis in Neutral Solution and the Transition State Affinities of Proteases". Journal of the American Chemical Society. 118 (26): 6105–6109. Bibcode:1996JAChS.118.6105R. doi:10.1021/ja954077c.
- ^ Keller J (2013). "Gastrointestinal Digestion and Absorption". Encyclopedia of Biological Chemistry. Elsevier. pp. 354–359. doi:10.1016/b978-0-12-378630-2.00106-7. ISBN 978-0-12-378631-9.
- ^ Oda K (2012). "New families of carboxyl peptidases: serine-carboxyl peptidases and glutamic peptidases". Journal of Biochemistry. 151 (1): 13–25. doi:10.1093/jb/mvr129. PMID 22016395.
- ^ Switzar L, Giera M, Niessen WM (1 March 2013). "Protein Digestion: An Overview of the Available Techniques and Recent Developments". Journal of Proteome Research. 12 (3): 1067–1077. doi:10.1021/pr301201x. PMID 23368288.
- ^ Drauz K, Grayson I, Kleemann A, Krimmer HP, Leuchtenberger W, Weckbecker C (2007). "Amino Acids". Ullmann's Encyclopedia of Industrial Chemistry. doi:10.1002/14356007.a02_057.pub2. ISBN 978-3-527-30385-4.
- ^ Gosline J, Lillie M, Carrington E, Guerette P, Ortlepp C, Savage K (February 2002). Bailey AJ, Macmillan J, Shrewry PR, Tatham AS (eds.). "Elastic proteins: biological roles and mechanical properties". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 357 (1418): 121–132. doi:10.1098/rstb.2001.1022. PMC 1692928. PMID 11911769.
- ^ Maruyama K, Natori R, Nonomura Y (July 1976). "New elastic protein from muscle". Nature. 262 (5563): 58–60. Bibcode:1976Natur.262...58M. doi:10.1038/262058a0. PMID 934326.
- ^ Tskhovrebova L, Trinick J (February 2012). "Making muscle elastic: the structural basis of myomesin stretching". PLOS Biology. 10 (2): e1001264. doi:10.1371/journal.pbio.1001264. PMC 3279349. PMID 22347814.
- ^ Mizraji E, Acerenza L, Lin J (November 1987). "Viscoelastic models for enzymes with multiple conformational states". Journal of Theoretical Biology. 129 (2): 163–175. Bibcode:1987JThBi.129..163M. doi:10.1016/s0022-5193(87)80010-3. PMID 3455460.
- ^ Schiller T, Scheibel T (2024-04-18). "Bioinspired and biomimetic protein-based fibers and their applications". Communications Materials. 5 (1) 56. Bibcode:2024CoMat...5...56S. doi:10.1038/s43246-024-00488-2.
- ^ Sun J, He H, Zhao K, Cheng W, Li Y, Zhang P, et al. (September 2023). "Protein fibers with self-recoverable mechanical properties via dynamic imine chemistry". Nature Communications. 14 (1) 5348. Bibcode:2023NatCo..14.5348S. doi:10.1038/s41467-023-41084-1. PMC 10475138. PMID 37660126.
- ^ a b c d e Guthold M, Liu W, Sparks EA, Jawerth LM, Peng L, Falvo M, et al. (2007-10-02). "A comparison of the mechanical and structural properties of fibrin fibers with other protein fibers". Cell Biochemistry and Biophysics. 49 (3): 165–181. doi:10.1007/s12013-007-9001-4. PMC 3010386. PMID 17952642.
- ^ Wang K, Meng X, Guo Z (2021). "Elastin Structure, Synthesis, Regulatory Mechanism and Relationship With Cardiovascular Diseases". Frontiers in Cell and Developmental Biology. 9: 596702. doi:10.3389/fcell.2021.596702. PMC 8670233. PMID 34917605.
- ^ Debelle L, Tamburro AM (February 1999). "Elastin: molecular description and function". The International Journal of Biochemistry & Cell Biology. 31 (2): 261–272. doi:10.1016/S1357-2725(98)00098-3. PMID 10216959.
- ^ a b Khoury LR, Popa I (November 2019). "Chemical unfolding of protein domains induces shape change in programmed protein hydrogels". Nature Communications. 10 (1) 5439. Bibcode:2019NatCo..10.5439K. doi:10.1038/s41467-019-13312-0. PMC 6884551. PMID 31784506.
- ^ Tan R, Shin J, Heo J, Cole BD, Hong J, Jang Y (October 2020). "Tuning the Structural Integrity and Mechanical Properties of Globular Protein Vesicles by Blending Crosslinkable and NonCrosslinkable Building Blocks". Biomacromolecules. 21 (10): 4336–4344. doi:10.1021/acs.biomac.0c01147. PMID 32955862.
- ^ Souza PC, Alessandri R, Barnoud J, Thallmair S, Faustino I, Grünewald F, et al. (April 2021). "Martini 3: a general purpose force field for coarse-grained molecular dynamics" (PDF). Nature Methods. 18 (4): 382–388. doi:10.1038/s41592-021-01098-3. PMID 33782607.
- ^ "Piotr Szymczak's Homepage". www.fuw.edu.pl. Retrieved 2024-05-13.
- ^ Mapplebeck S, Booth J, Shalashilin D (August 2021). "Simulation of protein pulling dynamics on second time scale with boxed molecular dynamics". The Journal of Chemical Physics. 155 (8): 085101. Bibcode:2021JChPh.155h5101M. doi:10.1063/5.0059321. PMID 34470356.
- ^ Carrion-Vazquez M, Marszalek PE, Oberhauser AF, Fernandez JM (September 1999). "Atomic force microscopy captures length phenotypes in single proteins". Proceedings of the National Academy of Sciences of the United States of America. 96 (20): 11288–11292. Bibcode:1999PNAS...9611288C. doi:10.1073/pnas.96.20.11288. PMC 18026. PMID 10500169.
- ^ Weinreb E, McBride JM, Siek M, Rougemont J, Renault R, Peleg Y, et al. (2025-03-28). "Enzymes as viscoelastic catalytic machines". Nature Physics. 21 (5): 787–798. Bibcode:2025NatPh..21..787W. doi:10.1038/s41567-025-02825-9. ISSN 1745-2481.
- ^ McKittrick J, Chen PY, Bodde SG, Yang W, Novitskaya EE, Meyers MA (2012-04-03). "The Structure, Functions, and Mechanical Properties of Keratin". JOM. 64 (4): 449–468. Bibcode:2012JOM....64d.449M. doi:10.1007/s11837-012-0302-8.
- ^ Yang L, van der Werf KO, Fitié CF, Bennink ML, Dijkstra PJ, Feijen J (March 2008). "Mechanical properties of native and cross-linked type I collagen fibrils". Biophysical Journal. 94 (6): 2204–2211. Bibcode:2008BpJ....94.2204Y. doi:10.1529/biophysj.107.111013. PMC 2257912. PMID 18032556.
- ^ Lessen HJ, Fleming PJ, Fleming KG, Sodt AJ (August 2018). "Building Blocks of the Outer Membrane: Calculating a General Elastic Energy Model for β-Barrel Membrane Proteins". Journal of Chemical Theory and Computation. 14 (8): 4487–4497. doi:10.1021/acs.jctc.8b00377. PMC 6191857. PMID 29979594.
Further reading
[edit]- Textbooks
- Branden C, Tooze J (1999). Introduction to Protein Structure. New York: Garland Pub. ISBN 978-0-8153-2305-1.
- Murray RF, Harper HW, Granner DK, Mayes PA, Rodwell VW (2006). Harper's Illustrated Biochemistry. New York: Lange Medical Books/McGraw-Hill. ISBN 978-0-07-146197-9.
- Van Holde KE, Mathews CK (1996). Biochemistry. Menlo Park, California: Benjamin/Cummings Pub. Co., Inc. ISBN 978-0-8053-3931-4.
- History
- Tanford C, Reynolds JA (2001). Nature's Robots: A History of Proteins. Oxford New York: Oxford University Press, USA. ISBN 978-0-19-850466-5.
External links
[edit]Databases and projects
[edit]- NCBI Entrez Protein database
- NCBI Protein Structure database
- Human Protein Reference Database
- Human Proteinpedia
- Folding@Home (Stanford University) Archived 2012-09-08 at the Wayback Machine
- Protein Databank in Europe (see also PDBeQuips, short articles and tutorials on interesting PDB structures)
- Research Collaboratory for Structural Bioinformatics (see also Molecule of the Month Archived 2020-07-24 at the Wayback Machine, presenting short accounts on selected proteins from the PDB)
- Proteopedia – Life in 3D: rotatable, zoomable 3D model with wiki annotations for every known protein molecular structure.
- UniProt the Universal Protein Resource
Tutorials and educational websites
[edit]- "An Introduction to Proteins" from HOPES (Huntington's Disease Outreach Project for Education at Stanford)
- Proteins: Biogenesis to Degradation – The Virtual Library of Biochemistry and Cell Biology
 KSF
KSF